Methanogens
Archaea · other
Methanogenic archaea
Name meaning: "From Greek 'methane' (methane) and '-genes' (producing), meaning 'methane-producing archaea'."
Discovery
Microorganism traits
Habitat
Classification history
1936: Kluyver & van Niel named the genera Methanobacterium and Methanosarcina. 1977: Woese & Fox demonstrated via 16S rRNA analysis that methanogens belong to Archaea, a domain distinct from Bacteria. Traditionally placed in phylum Euryarchaeota; GTDB reclassifies them into Methanobacteriota, Halobacteriota, and Thermoplasmatota. Metagenomics has revealed MCR-encoding lineages in Thermoproteota and Asgardarchaeota.
Clinical significance
No direct pathogenicity reported. Human gut commensal (*Methanobrevibacter smithii*) involved in digestive efficiency; excessive methane production may be associated with constipation, bloating, and IBS-C.

Methanogens (methanogenic archaea) are a polyphyletic assemblage of archaea that produce methane (CH₄) as the obligate end product of their energy metabolism. Belonging to the domain Archaea, they are strictly anaerobic organisms that carry out methanogenesis — the biological production of methane — as their sole means of generating ATP. Substrates include hydrogen and carbon dioxide, acetate, methanol, methylamines, and other methylated compounds. All methanogens share one defining biochemical feature: the methyl-coenzyme M reductase (MCR) enzyme system, which catalyzes the terminal step in methane formation.
Methanogens are the only biological source of methane on Earth, accounting for approximately 60–70% of total global methane emissions. They inhabit a remarkably diverse range of anaerobic environments — wetlands, marine sediments, the digestive tracts of ruminants, termite hindguts, the human intestine, wastewater treatment plants, rice paddies, landfills, and deep-sea hydrothermal vents. Over 150 species have been cultivated and formally described (DSMZ), distributed across at least three phyla and more than seven orders, confirming their status as a polyphyletic group united by shared metabolism rather than shared ancestry. Genome sizes span from approximately 1.24 Mb (Methanothermus fervidus) to 5.75 Mb (Methanosarcina acetivorans).
Methanogens are among the oldest life forms on Earth, with geochemical and molecular clock evidence suggesting that methanogenesis originated approximately 3.5 billion years ago. They play a central role in the global carbon cycle and climate system: methane is a potent greenhouse gas with a 100-year global warming potential (GWP) of 28–34 relative to CO₂. At the same time, the metabolic activity of methanogens is harnessed industrially for biogas production, a significant renewable energy source. In 1936, A. J. Kluyver and C. B. van Niel in the Netherlands first formally named methanogenic genera (Methanobacterium, Methanosarcina), and in the same year H. A. Barker reported the first cultivation of a methanogen (Methanobacterium omelianskii, later found to be a syntrophic co-culture), inaugurating the modern era of methanogen research.
1. Overview
The name "methanogen" derives from the Greek methane and -genes ("producing"), directly reflecting the defining metabolic characteristic of these organisms: the obligate production of methane gas as a byproduct of energy conservation.
Taxonomically, methanogens do not constitute a monophyletic group. They are a polyphyletic assemblage — a functional group united by shared metabolism (methanogenesis) rather than by descent from a single common ancestor. Traditionally classified within the phylum Euryarchaeota, methanogens have been redistributed under the GTDB (Genome Taxonomy Database) classification into the phyla Methanobacteriota, Halobacteriota, and Thermoplasmatota. Recent metagenomic studies have further expanded the known phylogenetic distribution of MCR-encoding genes to lineages within Thermoproteota (formerly the TACK superphylum) and Asgardarchaeota, revealing that the capacity for methane metabolism is far more widespread across the archaeal tree than previously appreciated.
The single most important shared characteristic of all methanogens is their strict anaerobiosis. Exposure to oxygen causes rapid, irreversible damage to key enzymes — particularly MCR and its nickel-containing cofactor F₄₃₀ — leading to cell death within minutes. This extreme oxygen sensitivity necessitates specialized anaerobic techniques for cultivation and study.
2. Taxonomy and Phylogeny
All cultivated methanogens belong to the domain Archaea. Under the traditional classification system, they were placed within the phylum Euryarchaeota. However, the GTDB system (release 95 and later), based on whole-genome phylogenetic analyses, splits the former Euryarchaeota into multiple phyla: methanogens are now distributed across Methanobacteriota, Halobacteriota, and Thermoplasmatota.
Within the traditional framework, methanogens are divided into five to seven orders. Methanobacteriales comprises rod-shaped, hydrogenotrophic methanogens including the genera Methanobacterium, Methanobrevibacter, and Methanothermobacter; notably, Methanobrevibacter smithii, the dominant archaeon of the human gut, belongs to this order. Methanococcales contains coccoid, marine methanogens frequently found at hydrothermal vents. Methanomicrobiales exhibits diverse cell morphologies and is common in freshwater and marine sediments. Methanosarcinales is the most metabolically versatile group, capable of utilizing hydrogen, acetate, and methylated compounds; its key genera include Methanosarcina and Methanotrix (formerly Methanosaeta). Methanopyrales is represented by a single hyperthermophilic genus, Methanopyrus, whose type species M. kandleri can grow at 122°C.
More recently recognized orders include Methanocellales (first discovered in rice paddy soils) and Methanomassiliicoccales (found in animal guts, using methylated compounds with hydrogen as an electron donor). Methanomassiliicoccales is classified within the phylum Thermoplasmatota under GTDB, phylogenetically distant from the traditional Euryarchaeota-associated methanogens.
The evolutionary origin of methanogenesis remains actively debated. A 2023 study in PNAS Nexus (Adam et al.) presented evidence that methane and alkane metabolism may have preceded the origin of Archaea themselves. Some researchers support the hypothesis that the Last Archaeal Common Ancestor (LACA) was a methanogen, while others argue that methanogenesis was horizontally transferred among multiple archaeal lineages. The distribution pattern of the MCR gene cluster (mcrABCDG) across disparate lineages is best explained by a combination of vertical inheritance and horizontal gene transfer.
| Taxonomic Level | Traditional Classification | GTDB Classification |
|---|---|---|
| Domain | Archaea | Archaea |
| Phylum | Euryarchaeota | Methanobacteriota, Halobacteriota, Thermoplasmatota |
| Representative Classes | Methanobacteria, Methanococci, Methanopyri, Methanomicrobia | Methanobacteria, Methanococci, Methanopyri, Methanocellia, etc. |
| Representative Orders | Methanobacteriales, Methanococcales, Methanomicrobiales, Methanosarcinales, Methanopyrales, Methanocellales, Methanomassiliicoccales | (same) |
3. Morphology and Structure
Methanogens display remarkable morphological diversity. Coccoid cells are found in Methanococcus and Methanothermococcus (diameter ~0.5–2 μm). Rod-shaped cells characterize Methanobacterium and Methanothermobacter (length ~2–10 μm, width ~0.4–1 μm). Methanospirillum hungatei exhibits a spiral morphology (~7 μm long). Methanosarcina forms irregular coccoid cells arranged in large aggregates, where individual cells measure ~1–3 μm but clusters may reach tens of micrometers. Plate-shaped cells are found in Methanoplanus.
The cell wall architecture of methanogens differs fundamentally from the peptidoglycan of bacteria. Species in Methanobacteriales and Methanopyrales possess pseudopeptidoglycan, which is structurally reminiscent of bacterial peptidoglycan but chemically distinct, containing N-acetyltalosaminuronic acid and β(1→3) glycosidic bonds. Most methanogens have an S-layer (surface layer) composed of protein or glycoprotein subunits arranged in a crystalline lattice on the cell surface. Certain Methanosarcina species produce a unique extracellular polysaccharide called methanochondroitin.
The cell membrane is composed of isoprenoid ether lipids characteristic of archaea. Unlike the ester-linked fatty acid lipids of bacteria and eukaryotes, the ether linkages (-O-) in archaeal lipids confer enhanced stability under extreme conditions. Some thermophilic species possess tetraether lipids that span the entire membrane, forming a monolayer that increases thermal stability.
Some methanogens possess archaella (singular: archaellum), the archaeal equivalent of bacterial flagella. Although functionally analogous, archaella differ in structure and assembly mechanism; their filament diameter (~10–13 nm) is thinner than that of bacterial flagella (~20 nm). Certain species also possess gas vesicles for buoyancy regulation.
The most distinctive visual marker of methanogens is F₄₂₀ autofluorescence. The presence of coenzyme F₄₂₀ causes cells to emit blue-green fluorescence (~470 nm) when excited with UV light at 420 nm. This property is routinely used to identify and quantify methanogens in complex microbial communities using epifluorescence microscopy.
4. Genome and Molecular Biology
Methanogen genomes consist of circular double-stranded DNA, with sizes varying widely across species. A comparative genomic study of the methanogen core and pangenome (Prondzinsky et al., 2023) found that genome sizes among cultivated methanogens range from approximately 1.24 Mb (Methanothermus fervidus) to 5.75 Mb (Methanosarcina acetivorans). GC content spans ~27–62%, and most species encode 1,500–3,500 protein-coding genes. Thermotolerant methanogens tend to have genomes roughly 30% smaller on average than psychrotolerant species, suggesting that temperature exerts a stronger influence on genome size than phylogeny.
Analysis of 86 cultivated methanogens revealed an extended core genome of 552 orthogroups, representing approximately 33.7% of a given genome. The majority of core genes (57%) are involved in biosynthesis, followed by coenzyme transport and metabolism (13%). The retention of such a large core genome across three phyla underscores the strong constraint that shared methanogenic biochemistry imposes on genome content.
Key gene clusters essential for methanogenesis include mcrABCDG, encoding the methyl-coenzyme M reductase that catalyzes the terminal methane-forming step; fmdABC/fwdABC, encoding formylmethanofuran dehydrogenase for the initial CO₂ reduction step; and mtrA-H, encoding the methyl-H₄MPT:CoM methyltransferase responsible for energy-conserving sodium ion translocation. Among these, mcrA is universally conserved across all methanogens and serves as the standard molecular marker for detection and diversity analysis.
Regarding thermal adaptation, thermotolerant methanogens specifically harbor reverse gyrase and metal transporter genes, while psychrotolerant species are enriched in genes related to cell membrane and wall biosynthesis, chaperones, and motility. Amino acid composition analysis reveals that thermophilic species show increased charged amino acids (lysine, glutamic acid) and hydrophobic residues (leucine, isoleucine), while psychrotolerant species show increased polar uncharged amino acids (glutamine, threonine).
Horizontal gene transfer (HGT) has played a significant role in methanogen evolution. Evidence for independent transfer of methanogenesis-related gene clusters across multiple lineages has been reported, and this is one proposed explanation for the polyphyletic distribution of methanogens.
5. Biochemistry and Metabolism
Methanogenesis proceeds through three major pathways. The hydrogenotrophic pathway is the most widespread: 4H₂ + CO₂ → CH₄ + 2H₂O (ΔG°' = −131 kJ/mol). The acetoclastic pathway cleaves acetate: CH₃COOH → CH₄ + CO₂ (ΔG°' = −36 kJ/mol); only Methanosarcina and Methanotrix can perform this reaction. The methylotrophic pathway uses methylated compounds such as methanol, methylamines, and dimethyl sulfide as substrates.
All three pathways converge at the terminal step catalyzed by MCR: CH₃-S-CoM + HS-CoB → CH₄ + CoM-S-S-CoB (ΔG°' ≈ −30 kJ/mol).
Methanogens employ a suite of unique coenzymes and cofactors not found in other organisms. Coenzyme M (2-mercaptoethanesulfonic acid) is the smallest known organic coenzyme in nature, serving as the methyl carrier in the final step. Coenzyme B (7-mercaptoheptanoylthreonine phosphate) participates alongside CoM in methane formation. Coenzyme F₄₂₀, a 5-deazaflavin derivative, functions as an electron carrier and is responsible for the characteristic fluorescence. Coenzyme F₄₃₀, a nickel-containing tetrapyrrole (hydrocorphinoid), is the prosthetic group of MCR and directly catalyzes methane formation; its structural elucidation required over 30 years of research. Methanofuran is the initial CO₂-binding cofactor, and tetrahydromethanopterin (H₄MPT) serves as the C₁ carrier during the reduction sequence.
Energy conservation mechanisms include chemiosmotic coupling (Na⁺/H⁺ translocation by the Mtr methyltransferase), electron bifurcation, and heterodisulfide reductase (Hdr)-mediated energy conservation. Given the thermodynamically marginal energy yields of methanogenesis, these diverse strategies are essential for the viability of methanogenic metabolism.
6. Ecology and Environmental Role
Methanogens are central players in the global carbon cycle, performing the terminal step of organic matter degradation in anaerobic environments. Complex organic matter is progressively broken down through successive fermentation stages to hydrogen, carbon dioxide, and acetate, which methanogens then convert to methane, completing the mineralization process.
Global methane emissions total approximately 580–610 Tg (teragrams) per year (Global Methane Budget 2024), of which roughly 60–70% is of biological origin via methanogenesis. Natural wetlands represent the single largest natural source at 150–220 Tg/yr. Ruminant livestock (primarily cattle) contribute approximately 90–100 Tg/yr, rice paddies 25–50 Tg/yr, landfills 50–70 Tg/yr, and termites 10–20 Tg/yr.
One of the most ecologically critical functions of methanogens is interspecies hydrogen transfer. By consuming hydrogen produced during fermentation, methanogens maintain low hydrogen partial pressures, thereby making otherwise thermodynamically unfavorable fermentation reactions possible. This syntrophic relationship is indispensable: without methanogens, the entire anaerobic degradation chain would stall.
In the rumen of ruminant animals, methanogens form complex microbial communities with cellulolytic bacteria and protozoa. A single cow emits approximately 70–120 L of methane per day, predominantly through eructation (belching; >90% of emissions), representing a loss of approximately 2–12% of ingested feed energy. In termite hindguts, methanogens live in symbiosis with cellulose-degrading protozoa, and some methanogens have been found as endosymbionts within the protist cells themselves.
In marine sediments, close relatives of methanogens — anaerobic methanotrophic archaea (ANME) — perform reverse methanogenesis, oxidizing methane using sulfate as the terminal electron acceptor. ANME consortia remove approximately 80–90% of methane produced in marine sediments before it reaches the atmosphere, consuming an estimated 200–300 Tg of methane annually and functioning as a critical methane sink.
7. Climate Change and Methane Mitigation
Methane is the second most important anthropogenic greenhouse gas after carbon dioxide. Its 100-year GWP is approximately 28–34, and its 20-year GWP is approximately 80–86, making it far more potent per unit mass than CO₂ over shorter timescales. According to NOAA data, atmospheric methane concentrations have risen from approximately 700 ppb in the pre-industrial era to approximately 1,922–1,930 ppb as of 2024, contributing an estimated 30% of total anthropogenic global warming.
Intensive research is underway to reduce methane emissions from ruminant livestock. 3-Nitrooxypropanol (3-NOP, marketed as Bovaer), which binds to the MCR cofactor F₄₃₀ and specifically inhibits methanogenesis, received FDA approval in the United States in 2024 for use in dairy cattle. Meta-analyses indicate an average methane reduction of approximately 30–33% in dairy cattle, with reductions of up to 59% under certain conditions. Other approaches under investigation include macroalgae (Asparagopsis taxiformis), tannins, saponins, anti-methanogen vaccines, and selective breeding for low-methane-emitting livestock.
Climate change itself may create positive feedback loops that amplify methane emissions. Thawing of Arctic permafrost exposes previously frozen organic matter to microbial decomposition, potentially releasing large quantities of methane. Rising temperatures may expand wetland areas and accelerate methanogenic activity. Destabilization of submarine methane hydrates is a longer-term concern, though the timescale and magnitude remain uncertain.
8. Industrial Applications and Biogas
Methanogens are the key microorganisms in anaerobic digestion, the process by which organic waste is converted to biogas. In wastewater treatment plants, anaerobic digestion reduces sludge volume by 40–60% while recovering energy in the form of biogas composed of approximately 60–70% methane and 30–40% carbon dioxide. Large-scale facilities can generate enough biogas to offset a substantial fraction — in some cases up to 100% — of their electricity consumption.
Optimal operating conditions for anaerobic digesters include mesophilic (35–40°C) or thermophilic (50–60°C) temperature ranges, pH 6.8–7.5, and a hydraulic retention time (HRT) of 15–30 days. Organic overloading leads to volatile fatty acid accumulation that inhibits methanogenesis, making careful process control essential.
A promising emerging technology is biomethanation, a Power-to-Gas approach in which electrolytically produced hydrogen and industrial CO₂ are converted to methane by methanogens. This represents a potential pathway for renewable energy storage, converting intermittent electricity into storable, pipeline-compatible methane.
9. Relationship to Human Health
The most common methanogen in the human gut is Methanobrevibacter smithii, detectable in approximately 30–96% of adults. By consuming hydrogen generated by fermentative gut bacteria, M. smithii facilitates more efficient polysaccharide fermentation, enabling greater energy extraction from dietary fiber.
Excessive intestinal methane production has been associated with constipation, bloating, and other gastrointestinal symptoms. Methane is known to slow intestinal transit, and breath methane testing is used in the diagnosis of methane-dominant irritable bowel syndrome with constipation (IBS-C) and small intestinal bacterial overgrowth (SIBO). A fasting breath methane concentration above 10 ppm is considered indicative of active methanogenic colonization. Some studies have reported a positive correlation between M. smithii abundance and obesity, but results are inconsistent, and a causal relationship has not been established.
No methanogen has been demonstrated to be directly pathogenic to humans. They are classified at BSL-1 and are not associated with infectious disease.
10. Extremophilic Adaptations
Methanogens collectively span one of the widest growth temperature ranges of any single physiological group in biology. Methanococcoides burtonii, isolated from Ace Lake in Antarctica, grows at temperatures as low as −2.5°C, while Methanopyrus kandleri strain 116 can proliferate at 122°C under high pressure (20 MPa) — the highest growth temperature recorded for any organism.
The hyperthermophilic adaptation of M. kandleri involves accumulation of intracellular cations (K⁺, Mg²⁺), a unique protective solute (cyclic 2,3-diphosphoglycerate) for protein stabilization, and post-transcriptional modification of tRNA to enhance nucleic acid thermostability. Intriguingly, proteome amino acid composition analysis reveals that M. kandleri resembles mesophilic rather than thermophilic methanogens in many properties (bulkiness, flexibility, solvent-accessible surface area), suggesting the existence of as-yet-undiscovered thermoadaptation mechanisms.
Certain halophilic methanogens survive NaCl concentrations exceeding 5 M, and haloalkaliphilic species such as Methanosalsum zhilinae thrive at pH values above 10.
11. Research History and Scientific Significance
The recognition of biogenic methane dates to 1776, when Alessandro Volta identified "marsh gas" from lake sediments. However, the microbial origin of this gas was not scientifically established until the late 19th and early 20th centuries.
The year 1936 was a landmark in methanogen research. A. J. Kluyver and C. B. van Niel formally named the genera Methanobacterium and Methanosarcina, recognizing methanogens as a distinct metabolic group. In the same year, H. A. Barker reported the first cultivation of a methanogen, Methanobacterium omelianskii, which was later shown to be a syntrophic co-culture of two organisms rather than a pure culture.
In 1977, Carl Woese and George Fox used 16S rRNA sequence analysis to demonstrate that methanogens were fundamentally distinct from bacteria, providing the critical evidence for establishing Archaea as the third domain of life. This discovery is widely regarded as one of the most important advances in 20th-century biology, transforming our understanding of the tree of life.
Subsequently, R. S. Wolfe and W. E. Balch developed innovative anaerobic cultivation techniques in the late 1970s that enabled the isolation and characterization of numerous methanogen species. In 2008, Takai et al. reported the growth of M. kandleri strain 116 at 122°C, setting the current upper temperature limit for life.
Current research on methanogens spans climate change mitigation (reducing methane emissions), bioenergy (biogas and Power-to-Gas technologies), astrobiology (methanogens as model organisms for life detection on Mars and icy moons), and the gut microbiome (the role of methanogens in human health and disease). The detection of seasonal methane fluctuations in the Martian atmosphere has placed methanogens at the center of astrobiological investigations.
12. Comparison with Related Microorganisms
| Feature | Methanogens | Haloarchaea | ANME (Anaerobic Methanotrophs) | Aerobic Methanotrophs |
|---|---|---|---|---|
| Energy metabolism | Methane production | Respiration/fermentation/phototrophy | Methane oxidation (reverse methanogenesis) | Methane oxidation (aerobic) |
| Classification | Archaea (polyphyletic) | Archaea (Halobacteriota) | Archaea (Halobacteriota, etc.) | Bacteria (Proteobacteria, etc.) |
| Oxygen requirement | Strictly anaerobic | Aerobic/facultative | Strictly anaerobic | Aerobic/microaerophilic |
| Key enzyme | MCR (forward) | Bacteriorhodopsin | MCR (reverse) | Methane monooxygenase (MMO) |
| Ecological role | Carbon mineralization; methane source | Decomposers in hypersaline habitats | Marine methane sink | Atmospheric/soil methane sink |
| Habitat | Diverse anaerobic environments | Hypersaline environments | Marine sediments | Soils, wetlands, freshwater |
Methanogens and ANME share the MCR enzyme, but ANME operate it in reverse to oxidize methane. The two groups are phylogenetically related, and ANME represent an evolutionary exploitation of the reversibility of the methanogenesis pathway.
13. Unresolved Questions
Several major questions remain open in methanogen research. The precise evolutionary origin of methanogenesis — whether it was present in the Last Archaeal Common Ancestor (LACA) or was subsequently acquired via horizontal gene transfer — is still debated. The evolutionary relationship between DPANN archaea and methanogens is unclear. The magnitude and timescale of methane release from thawing permafrost remain a major uncertainty in climate modeling. The causal relationship between gut methanogens and obesity, metabolic disorders, and digestive diseases has not been established. The true diversity and ecological functions of uncultivated environmental methanogens are only beginning to be revealed through metagenomics. Developing effective strategies to reduce ruminant methane emissions while maintaining animal health and productivity continues to be an active area of research.
14. References
Kluyver, A.J. & van Niel, C.B. (1936). Prospects for a natural system of classification of bacteria. Zentralbl. Bakteriol. Parasitenkd. Infektionskr. Hyg. Abt. II, 94, 369–403.
Woese, C.R. & Fox, G.E. (1977). Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proceedings of the National Academy of Sciences, 74(11), 5088–5090. https://doi.org/10.1073/pnas.74.11.5088
Thauer, R.K., Kaster, A.K., Seedorf, H., et al. (2008). Methanogenic archaea: ecologically relevant differences in energy conservation. Nature Reviews Microbiology, 6(8), 579–591. https://doi.org/10.1038/nrmicro1931
Liu, Y. & Whitman, W.B. (2008). Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Annals of the New York Academy of Sciences, 1125(1), 171–189. https://doi.org/10.1196/annals.1419.019
Takai, K., Nakamura, K., Toki, T., et al. (2008). Cell proliferation at 122°C and isotopically heavy CH₄ production by a hyperthermophilic methanogen under high-pressure cultivation. PNAS, 105(31), 10949–10954. https://doi.org/10.1073/pnas.0712334105
Saunois, M., Stavert, A.R., Poulter, B., et al. (2020). The global methane budget 2000–2017. Earth System Science Data, 12(3), 1561–1623. https://doi.org/10.5194/essd-12-1561-2020
Lyu, Z., Shao, N., Akinyemi, T., & Whitman, W.B. (2018). Methanogenesis. Current Biology, 28(13), R566–R571. https://doi.org/10.1016/j.cub.2018.05.021
Buan, N.R. (2018). Methanogens: pushing the boundaries of biology. Emerging Topics in Life Sciences, 2(4), 629–646. https://doi.org/10.1042/ETLS20180031
Conrad, R. (2020). Importance of hydrogenotrophic, aceticlastic and methylotrophic methanogenesis for methane production in terrestrial, aquatic and other anoxic environments. Pedosphere, 30(1), 25–39. https://doi.org/10.1016/S1002-0160(19)60823-1
Hook, S.E., Wright, A.D.G., & McBride, B.W. (2010). Methanogens: methane producers of the rumen and mitigation strategies. Archaea, 2010, 945785. https://doi.org/10.1155/2010/945785
Prondzinsky, P., Toyoda, S., & McGlynn, S.E. (2023). The methanogen core and pangenome: conservation and variability across biology's growth temperature extremes. Microbial Genomics, 9(1), dsac048. https://doi.org/10.1093/femsml/uqac008
Adam, P.S., Borrel, G., & Gribaldo, S. (2023). The origin and evolution of methanogenesis and Archaea are intertwined. PNAS Nexus, 2(2), pgad023. https://doi.org/10.1093/pnasnexus/pgad023
Garcia, J.L., Patel, B.K.C., & Ollivier, B. (2000). Taxonomic, phylogenetic, and ecological diversity of methanogenic Archaea. Anaerobe, 6(4), 205–226. https://doi.org/10.1006/anae.2000.0345
Enzmann, F., Mayer, F., Rother, M., & Holtmann, D. (2018). Methanogens: biochemical background and biotechnological applications. AMB Express, 8(1), 1–22. https://doi.org/10.1186/s13568-017-0531-x
Dridi, B., Henry, M., El Khéchine, A., Raoult, D., & Drancourt, M. (2009). High prevalence of Methanobrevibacter smithii and Methanosphaera stadtmanae detected in the human gut using an improved DNA detection protocol. PLoS ONE, 4(9), e7063. https://doi.org/10.1371/journal.pone.0007063
Hanišáková, N., Vítězová, M., et al. (2022). The historical development of cultivation techniques for methanogens and other strict anaerobes and their application in modern microbiology. Microorganisms, 10(2), 412. https://doi.org/10.3390/microorganisms10020412
Saunois, M., Stavert, A.R., Poulter, B., et al. (2025). Global Methane Budget 2000–2020. Earth System Science Data, 17, 1873–1958. https://doi.org/10.5194/essd-17-1873-2025
Adam, P.S., Borrel, G., Brochier-Armanet, C., & Gribaldo, S. (2017). The growing tree of Archaea: new perspectives on their diversity, evolution and ecology. The ISME Journal, 11(11), 2407–2425. https://doi.org/10.1038/ismej.2017.122
Fun Facts
Methanogens are among the oldest life forms on Earth, estimated to have existed since approximately 3.5 billion years ago. The oxygen-free atmosphere of early Earth was ideal for methanogenesis, and even after the Great Oxidation Event (~2.4 billion years ago) they survived by retreating into anaerobic niches where they persist to this day.
Methanopyrus kandleri strain 116 can grow at 122°C, holding the all-time record for the highest growth temperature of any known organism. This is above the normal boiling point of water and roughly the temperature inside a household pressure cooker (~120°C).
The world's cattle emit approximately 100 million tonnes (100 Tg) of methane per year — about 15–17% of all anthropogenic methane emissions. A single cow belches roughly 70–120 liters of methane daily, equivalent to the carbon emissions from driving a car approximately 30–50 km.
Methanogens glow blue-green under ultraviolet light, thanks to their unique coenzyme F₄₂₀. This natural fluorescence allows scientists to easily spot methanogens in complex microbial communities using an epifluorescence microscope — no staining required.
The discovery of methanogens changed biology forever. In 1977, Carl Woese's analysis of methanogen 16S rRNA revealed they were fundamentally different from all known bacteria, leading directly to the establishment of Archaea as the third domain of life — one of the most important scientific discoveries of the 20th century.
About 80–90% of methane produced in ocean sediments never reaches the atmosphere, thanks to ANME (anaerobic methanotrophic archaea) — close relatives of methanogens that run methanogenesis in reverse. This 'methane filter' removes an estimated 200–300 Tg of methane per year, preventing catastrophic additional greenhouse warming.
Coenzyme M (2-mercaptoethanesulfonic acid), used by all methanogens, is the smallest known organic coenzyme in nature. And coenzyme F₄₃₀, a nickel-containing molecule found exclusively in methanogens, took over 30 years of research to fully characterize structurally.
Methanobrevibacter smithii, the most common methanogen in the human gut, is found in approximately 30–96% of adults. By consuming hydrogen, it makes fiber digestion more efficient — but this also means more calories are extracted from food, prompting researchers to investigate a possible link to obesity.
Termites are tiny but mighty methane emitters: their hindgut methanogens contribute 10–20 Tg of methane annually, about 2–3% of global emissions. Some of these methanogens live not just in the termite gut, but actually inside the cells of gut-dwelling protists — a symbiosis within a symbiosis.
Large wastewater treatment plants can generate enough biogas from methanogen activity to power up to 100% of their own electricity needs — turning waste into energy while cleaning water. It's one of the most elegant circular economy technologies in use today.
Methanogens collectively span one of the widest growth temperature ranges of any physiological group in biology: from −2.5°C (in Antarctic lakes) to 122°C (at deep-sea hydrothermal vents) — a staggering range of 124.5°C.
FAQ
Methanogens are exclusively archaea, belonging to the domain Archaea. Although both bacteria and archaea are prokaryotes, they differ fundamentally in cell wall composition, membrane lipid chemistry (ether-linked vs. ester-linked), and genomic features. Notably, it was Carl Woese's analysis of methanogen 16S rRNA in 1977 that provided the key evidence for establishing Archaea as a separate domain of life. Methanogenesis — the biological production of methane — is found only in archaea and has never been observed in bacteria or eukaryotes.
Methanogens are strictly anaerobic and die within minutes of oxygen exposure. This is because their essential methanogenic enzymes — particularly the methyl-coenzyme M reductase (MCR) and its nickel-containing cofactor F₄₃₀ — are irreversibly oxidized and inactivated by molecular oxygen. Additionally, most methanogens lack or have very limited quantities of reactive oxygen species (ROS) detoxification enzymes such as catalase and superoxide dismutase, leaving them with virtually no defense against oxygen toxicity.
Contrary to popular belief, ruminant methane is overwhelmingly emitted via eructation (belching), not flatulence. Over 90% of methane is produced in the rumen and released through the mouth during the normal rumination process. Only about 10% or less is generated in the lower gastrointestinal tract and emitted as flatulence.
Yes, approximately 30–96% of adults (depending on the study) harbor methanogens in their intestines — primarily Methanobrevibacter smithii — and emit methane through breath and flatulence. However, there is substantial individual variation; some people have very few gut methanogens and produce negligible methane. A fasting breath methane level above 10 ppm is considered indicative of active methanogenic colonization, and this test is used clinically for diagnosing methane-dominant IBS-C and SIBO.
In certain sectors, yes — and practical progress is being made. The feed additive 3-NOP (marketed as Bovaer), which specifically inhibits the MCR enzyme, received FDA approval in 2024 for use in US dairy cattle and reduces enteric methane emissions by an average of ~30–33%, with some studies reporting up to 59%. However, suppressing methanogenesis in natural environments such as wetlands could have unpredictable ecological consequences and is not recommended. Climate mitigation efforts focus on controllable sources: livestock management, landfill gas capture, and rice paddy water management.
The phenomenon of biogenic methane was first recognized when Alessandro Volta identified 'marsh gas' from lake sediments in 1776. In 1936, Dutch microbiologists A. J. Kluyver and C. B. van Niel first formally named methanogenic genera (Methanobacterium, Methanosarcina), recognizing them as a distinct metabolic group. That same year, H. A. Barker reported the cultivation of Methanobacterium omelianskii (later found to be a co-culture). The decisive turning point came in 1977 when Carl Woese's 16S rRNA analysis revealed that methanogens belong to an entirely new domain of life — the Archaea — fundamentally distinct from bacteria.
Methanogens thrive in extreme environments (high temperature, high pressure, high salinity, high alkalinity), can generate energy from simple inorganic compounds (H₂ and CO₂), and require no oxygen — characteristics compatible with conditions on early Earth, Mars, and the subsurface oceans of icy moons. Seasonal methane fluctuations detected in the Martian atmosphere have prompted active investigation into whether methanogens or similar organisms could exist on Mars. Hypotheses for methanogen-like life beneath the surfaces of Jupiter's moon Europa and Saturn's moon Enceladus are also under study.
Both are primarily methane, but they differ in origin. Biogas is produced by methanogens actively decomposing organic matter in the present, containing approximately 50–70% methane, 30–40% CO₂, and trace amounts of H₂S. Natural gas was formed over millions to hundreds of millions of years through geological processes (some originating from ancient methanogenic activity) and is stored underground; it has a higher methane content (70–90%) and fewer impurities. From a carbon cycle perspective, biogas is considered renewable energy, while natural gas is a fossil fuel.
Methanopyrus kandleri strain 116 can grow at 122°C under high pressure (20 MPa), holding the record for the highest growth temperature of any known organism. High pressure raises water's boiling point, allowing the cell to function in a liquid environment. Protein stabilization involves intracellular cation accumulation (K⁺, Mg²⁺) and a unique protective solute (cyclic 2,3-diphosphoglycerate), while tRNA modifications enhance nucleic acid thermostability. Intriguingly, this organism's proteome amino acid composition resembles that of mesophilic rather than thermophilic methanogens, suggesting the existence of yet-undiscovered thermal adaptation mechanisms.
Yes — methanogens have a unique autofluorescence property due to their coenzyme F₄₂₀. When excited with UV light at 420 nm, methanogen cells emit blue-green fluorescence (~470 nm), making them readily distinguishable from other microorganisms under an epifluorescence microscope. This technique is practically used to identify and quantify methanogens in complex microbial communities such as anaerobic digester sludge and rumen contents. At the molecular level, PCR targeting the mcrA gene is the standard method for methanogen detection.
Methanogens perform the terminal step of the anaerobic organic matter degradation chain. Without them, hydrogen generated during fermentation would accumulate, raising hydrogen partial pressure to levels where upstream fermentation reactions become thermodynamically impossible. The entire anaerobic degradation process in wetlands, sediments, and animal guts would grind to a halt, profoundly disrupting nutrient cycling. Ruminant animals, in particular, would be unable to efficiently digest cellulose and would face severe nutritional consequences.
Gallery
4 images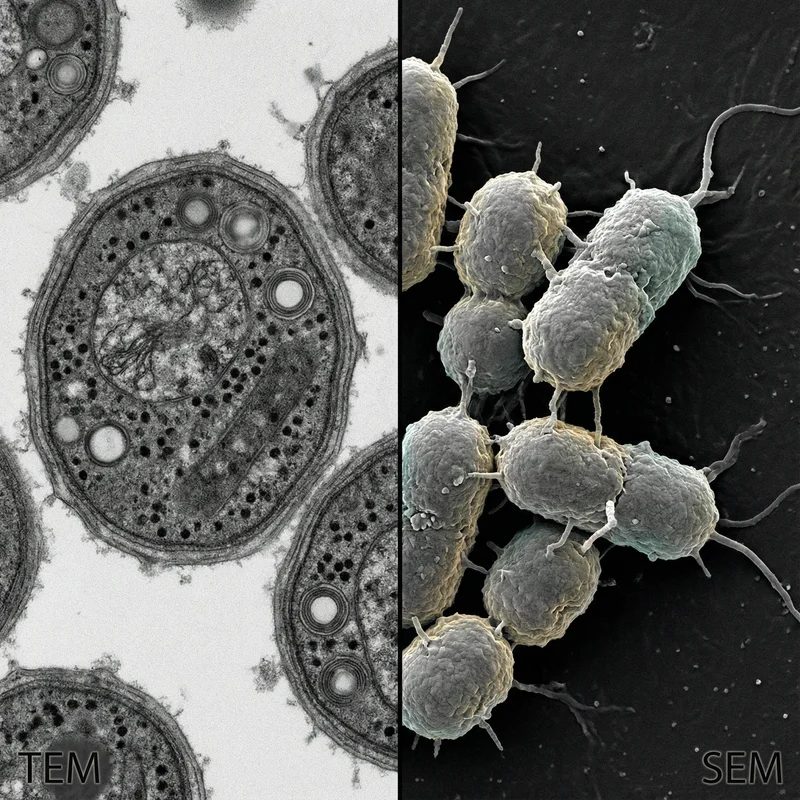 Methanogens
MethanogensMethanogens
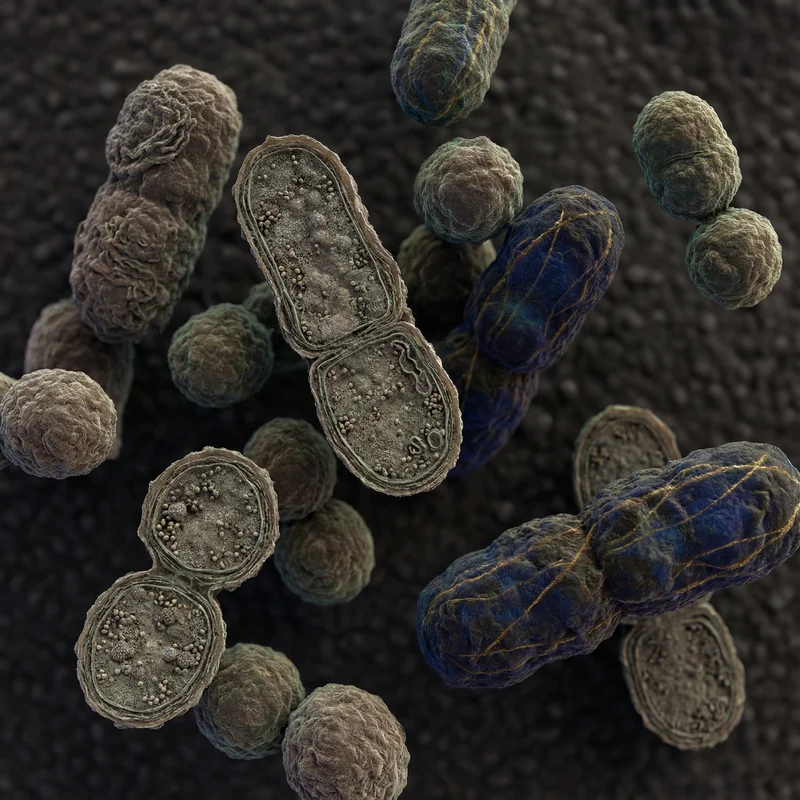 Methanogens
MethanogensMethanogens
 Methanogens
MethanogensMethanogens
 Methanogens
MethanogensMethanogens