Staphylococcus aureus
Bacteria · other
Staphylococcus aureus
Name meaning: "Grape cluster (staphyle, Greek) + grain/berry (kokkos, Greek) + golden (aureus, Latin)"
Discovery
Microorganism traits
Classification
Habitat
Classification history
1880: Alexander Ogston first observed staphylococci in surgical abscesses in Aberdeen, Scotland, and proposed the genus name Staphylococcus. 1884: Friedrich Julius Rosenbach formally named S. aureus based on golden colony pigmentation (originally as 'Staphylococcus pyogenes aureus'). Originally classified in Micrococcaceae; later reclassified to Staphylococcaceae based on molecular phylogenetics. 1980: Included on the Approved Lists of Bacterial Names. Recognized as the nomenclatural type species of genus Staphylococcus. Current correct name per LPSN; listed on LoRN for medically important bacteria
Clinical significance
Causes skin and soft tissue infections (SSTIs), bacteremia, sepsis, infective endocarditis, osteomyelitis, pneumonia, food poisoning, and toxic shock syndrome (TSS). MRSA is the deadliest pathogen-drug combination in global antimicrobial resistance, with approximately 130,000 directly attributable deaths in 2021. Listed as a WHO priority antibiotic-resistant pathogen
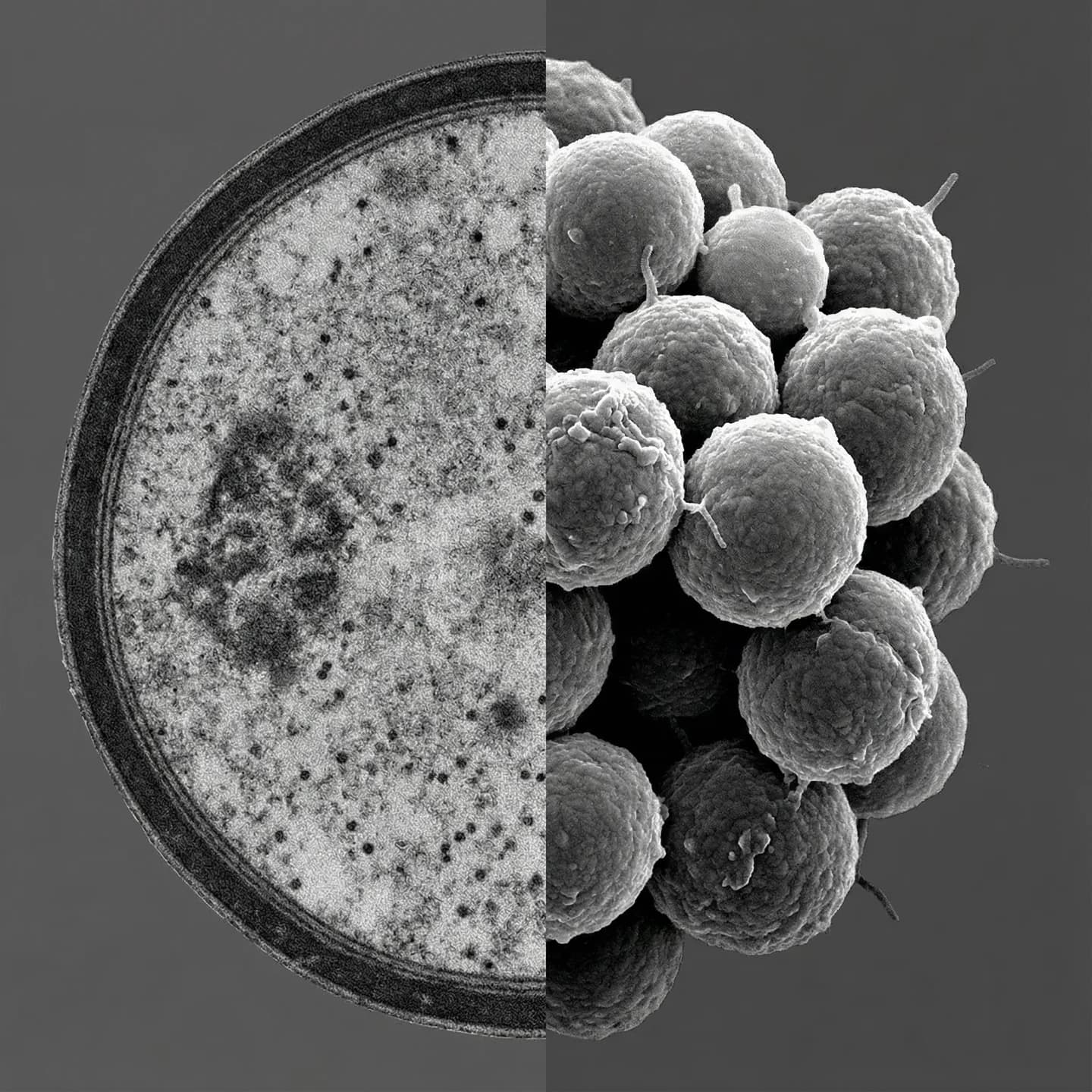
Staphylococcus aureus (Rosenbach, 1884) is a Gram-positive coccus belonging to the family Staphylococcaceae within the order Bacillales, and represents one of the most clinically significant human pathogens while simultaneously existing as a commensal member of the normal human microbiota. Cells are spherical, approximately 0.5 to 1.5 μm in diameter, and characteristically arrange in irregular grape-like clusters. When cultured on nutrient media, S. aureus produces distinctive golden-yellow colonies due to the carotenoid pigment staphyloxanthin. As a facultative anaerobe, it thrives in both aerobic and anaerobic environments. The organism is defined biochemically by being catalase-positive and coagulase-positive, the latter property being the key feature distinguishing it from most other staphylococcal species. Its genome consists of a single circular double-stranded DNA chromosome of approximately 2.7 to 3.1 Mb with a GC content of about 32 to 33%. It is classified as a Biosafety Level 2 (BSL-2) pathogen by the CDC and is assigned to Risk Group 2 by LPSN and international biosafety frameworks.
S. aureus persistently colonizes the anterior nares (nostrils) of approximately 20 to 30% of the global population, with another 30% being intermittent carriers (CDC, 2024). Under normal circumstances, carriage is asymptomatic, but when host defenses are compromised through skin breaks, surgical wounds, or immunosuppression, S. aureus can cause a vast spectrum of diseases ranging from minor skin infections to life-threatening invasive conditions such as bacteremia, sepsis, and endocarditis. This dual nature as both commensal and pathogen has earned it the designation of a "pathobiont." The World Health Organization (WHO) lists S. aureus among the most threatening antibiotic-resistant pathogens globally, and methicillin-resistant S. aureus (MRSA) in particular stands at the forefront of the antimicrobial resistance crisis.
According to the Global Burden of Disease (GBD) 2021 analysis published in The Lancet in 2024, deaths directly attributable to MRSA increased from approximately 57,200 in 1990 to 130,000 in 2021, while MRSA-associated deaths rose from 261,000 to 550,000 over the same period (GBD 2021 AMR Collaborators, 2024). In the United States alone, an estimated 120,000 S. aureus bloodstream infections occur annually, resulting in approximately 20,000 deaths (CDC, 2019). S. aureus bacteremia (SAB) has an estimated incidence of 20 to 30 cases per 100,000 person-years and a 30-day mortality of 15 to 40%, even with appropriate antimicrobial therapy (Tong et al., 2015; Nature, 2025). These figures underscore the enormous public health burden posed by this organism and its central role in modern infectious disease medicine and antimicrobial resistance research.
1. Overview
The genus name Staphylococcus derives from the Greek words 'σταφυλή' (staphylē, meaning grape cluster) and 'κόκκος' (kokkos, meaning grain or berry), reflecting the characteristic grape-like clustering of the cocci observed under microscopy. The species epithet aureus is the Latin word for "golden," referring to the golden-yellow pigmentation of colonies on culture media. This naming was established in 1884 by the German physician Friedrich Julius Rosenbach, who differentiated staphylococcal species by colony color, designating golden-colored colonies as S. aureus and white colonies as S. albus (now S. epidermidis).
In the LPSN (List of Prokaryotic names with Standing in Nomenclature), Staphylococcus aureus Rosenbach 1884 is recognized as a validly published and correct name. It is also the nomenclatural type species of the genus Staphylococcus and is listed on the List of Recommended Names for bacteria of medical importance (LoRN). According to NCBI Taxonomy, the full lineage is: Bacteria > Bacillota (Firmicutes) > Bacilli > Bacillales > Staphylococcaceae > Staphylococcus > S. aureus.
In a single sentence, S. aureus can be characterized as a Gram-positive, coagulase-positive coccus that is simultaneously one of the most common human commensals and one of the most lethal bacterial pathogens known.
2. Classification and Phylogeny
Taxonomic Position
The current taxonomic classification of S. aureus is as follows.
| Rank | Taxon |
|---|---|
| Domain | Bacteria |
| Phylum | Bacillota (synonym: Firmicutes) |
| Class | Bacilli |
| Order | Bacillales (synonym: Caryophanales) |
| Family | Staphylococcaceae |
| Genus | Staphylococcus |
| Species | S. aureus |
The genus Staphylococcus was originally classified within the family Micrococcaceae. However, molecular phylogenetic analyses revealed that staphylococci are not closely related to micrococci, and the genus was reclassified into the new family Staphylococcaceae (Schleifer and Bell, 2009). At the order level, NCBI Taxonomy places it under Bacillales, though some databases such as USDA ACIR use the synonym Caryophanales.
Naming History
In 1880, Scottish surgeon Alexander Ogston (1844-1929), working at Aberdeen Royal Infirmary, first observed grape-like clusters of bacteria in pus from surgical abscesses and proposed the genus name Staphylococcus. He presented his findings at the Ninth Surgical Congress in Berlin on April 9, 1880, demonstrating experimentally in mice that these bacteria were the causative agents of abscess formation. In 1884, Friedrich Julius Rosenbach formally described and named the species based on colony pigmentation, originally as "Staphylococcus pyogenes aureus," which was later simplified to S. aureus. The name was included on the Approved Lists of Bacterial Names (Skerman, McGowan, and Sneath, 1980), formally validating its nomenclatural standing.
Phylogenetic Analysis and Genetic Diversity
S. aureus strains are classified using a variety of molecular typing methods including multilocus sequence typing (MLST), spa typing, staphylococcal cassette chromosome mec (SCCmec) typing, and increasingly, whole-genome sequencing (WGS). MLST-based clonal complexes (CCs) define the major evolutionary lineages. Approximately 10 major human-associated lineages have been identified, including CC1, CC5, CC8, CC15, CC22, CC30, CC45, CC97, and CC398, each exhibiting distinct geographic distributions, host specificities, and resistance profiles.
CC8 encompasses USA300, the predominant CA-MRSA lineage in the United States, while CC22 (EMRSA-15) is the dominant HA-MRSA lineage in the United Kingdom and much of Europe. CC398 is primarily associated with livestock (especially swine) and represents the major livestock-associated MRSA (LA-MRSA) lineage, which can also transmit to humans in occupational settings.
Subspecies and Major Variants
Two validly published subspecies are recognized by LPSN: S. aureus subsp. aureus (the automatically created nominotypical subspecies) and S. aureus subsp. anaerobius (De La Fuente et al., 1985), the latter being a strict anaerobe that causes Morel disease (subcutaneous abscesses) in sheep.
Based on antibiotic susceptibility profiles, S. aureus strains are categorized as methicillin-susceptible S. aureus (MSSA), methicillin-resistant S. aureus (MRSA), vancomycin-intermediate S. aureus (VISA; vancomycin MIC 4-8 μg/mL), and vancomycin-resistant S. aureus (VRSA; vancomycin MIC ≥16 μg/mL). MRSA is further subdivided into healthcare-associated MRSA (HA-MRSA), community-associated MRSA (CA-MRSA), and livestock-associated MRSA (LA-MRSA), each with distinct epidemiological features, virulence factor profiles, and SCCmec types.
3. Morphology and Structure
Cell Morphology
S. aureus cells are spherical (cocci) with a diameter of approximately 0.5 to 1.5 μm. During cell division, the bacterium divides in two alternating perpendicular planes, and because daughter cells do not completely separate, they form the characteristic irregular grape-like clusters from which the genus name derives. Cells may also be observed as diplococci (pairs), short chains, or individually. S. aureus is nonmotile and does not form endospores.
Cell Wall Architecture
As a Gram-positive bacterium, S. aureus possesses a thick peptidoglycan layer (approximately 20-30 nm) that retains the crystal violet stain during Gram staining, rendering cells purple. The cell wall is rich in wall teichoic acid (WTA) and lipoteichoic acid (LTA), which play critical roles in cell wall homeostasis, autolysis regulation, antibiotic resistance, host cell adhesion, and biofilm formation. There is no outer membrane.
The cell surface displays an array of adhesins collectively known as MSCRAMMs (Microbial Surface Components Recognizing Adhesive Matrix Molecules). These proteins bind to host extracellular matrix components including fibronectin, fibrinogen, collagen, and elastin, and are essential for tissue colonization and infection initiation.
Pigmentation and Capsule
The golden color of S. aureus is produced by staphyloxanthin, a membrane-bound carotenoid pigment synthesized via the crtOPQMN gene cluster. Beyond its role in pigmentation, staphyloxanthin functions as a virulence factor by acting as an antioxidant that scavenges reactive oxygen species (ROS) and singlet oxygen generated by host neutrophils during the oxidative burst, thereby protecting the bacterium from killing by phagocytes. Mutant strains lacking staphyloxanthin are more readily killed by neutrophils, confirming the pigment's role in immune evasion.
Some strains produce a polysaccharide capsule, with capsular polysaccharide (CP) types 5 and 8 accounting for approximately 75 to 80% of clinical isolates. The capsule inhibits opsonization and phagocytosis, contributing to immune evasion.
4. Genome and Molecular Biology
Genome Structure
The S. aureus genome consists of a single circular double-stranded DNA chromosome. Based on numerous RefSeq entries in the NCBI database, genome sizes range from approximately 2.7 to 3.1 Mb, with reference strains such as NCTC 8325 having a genome of approximately 2.82 Mb. The GC content is relatively low at approximately 32 to 33%. The genome encodes an average of 2,500 to 2,800 protein-coding genes, of which approximately 56% constitute the core genome shared across all strains.
Mobile Genetic Elements
Approximately 15 to 20% of the S. aureus genome is composed of mobile genetic elements (MGEs), which represent the primary source of genetic diversity and adaptive potential. Key MGEs include prophages (bacteriophages integrated into the chromosome), staphylococcal pathogenicity islands (SaPIs), staphylococcal cassette chromosomes (SCCmec), insertion sequences (IS elements), transposons, and plasmids.
The SCCmec element is the central mobile genetic element harboring the methicillin resistance gene mecA (or its variant mecC). Over 14 SCCmec types have been identified to date. Types I, II, and III are primarily found in HA-MRSA strains and tend to carry additional resistance determinants, while types IV and V are typically associated with CA-MRSA strains and are smaller in size. SaPIs carry toxin genes such as those encoding TSST-1 and enterotoxins, and are mobilized via phage-mediated horizontal gene transfer.
Molecular Mechanisms of Antibiotic Resistance
S. aureus acquires antibiotic resistance through both mutation and horizontal gene transfer. Penicillin resistance, mediated by the blaZ-encoded beta-lactamase enzyme that hydrolyzes the beta-lactam ring, emerged within years of penicillin's clinical introduction in the 1940s and now affects over 90% of clinical isolates. Methicillin resistance is conferred by the mecA gene, which encodes an altered penicillin-binding protein (PBP2a/PBP2') with extremely low affinity for beta-lactam antibiotics, allowing continued cell wall synthesis in their presence.
Vancomycin-intermediate resistance (VISA) develops through stepwise mutations leading to thickened cell walls with increased D-Ala-D-Ala residues that sequester vancomycin before it can reach its target. Full vancomycin resistance (VRSA) results from acquisition of the vanA gene cluster from enterococci via plasmid-mediated conjugation, which alters the cell wall precursor terminus from D-Ala-D-Ala to D-Ala-D-Lac, reducing vancomycin binding affinity by over 1,000-fold. The first VRSA was isolated in 2002 in Michigan, United States.
A 2022 study published in Nature (Eiamphungporn et al.) revealed that the origin of mecC-mediated methicillin resistance predates the clinical use of antibiotics, tracing back to natural selection on the skin of European hedgehogs coinfected with S. aureus and the dermatophyte fungus Trichophyton erinacei, which produces the beta-lactam antibiotic trimethyldihydronapthoquinone. This finding demonstrates that antibiotic resistance can arise through natural ecological pressures independent of human antibiotic use.
5. Pathogenesis and Clinical Manifestations
Infection Mechanism
The pathogenesis of S. aureus infection follows a general sequence of adhesion, invasion, immune evasion, and tissue damage. During adhesion, MSCRAMMs (including fibronectin-binding proteins FnBPA/FnBPB, clumping factors ClfA/ClfB, and collagen-binding protein Cna) mediate attachment to host extracellular matrix proteins. During invasion, fibronectin-binding proteins trigger internalization into nonprofessional phagocytes such as epithelial and endothelial cells via integrin alpha-5/beta-1. Immune evasion involves a sophisticated arsenal including Protein A, capsular polysaccharides, leukocidins, complement inhibitory proteins (SCIN, Efb, Ecb), and superantigens.
Toxins and Enzymes
S. aureus secretes a broad array of toxins that damage host tissues. Pore-forming toxins include alpha-hemolysin (Hla/alpha-toxin), which oligomerizes into a heptameric pore in host cell membranes; bicomponent leukocidins such as Panton-Valentine leucocidin (PVL), gamma-hemolysin (HlgAB/HlgCB), LukED, and LukAB, which target and lyse leukocytes; and phenol-soluble modulins (PSMs), amphipathic alpha-helical peptides that disrupt membranes and also contribute to biofilm structuring and bacterial dissemination.
Superantigens include staphylococcal enterotoxins (over 20 serotypes: SEA through SEE and beyond) and toxic shock syndrome toxin-1 (TSST-1). These molecules crosslink the T-cell receptor (TCR) V-beta domain with MHC class II molecules on antigen-presenting cells, triggering nonspecific massive T-cell activation and a cytokine storm that can cause systemic inflammatory response and multi-organ failure. Exfoliative toxins (ETA and ETB) are serine proteases that cleave desmoglein 1 in the granular layer of the epidermis, causing the epidermal splitting characteristic of staphylococcal scalded skin syndrome (SSSS).
Key enzymes include coagulase and von Willebrand factor-binding protein (vWbp), which activate prothrombin to form fibrin clots that shield bacteria from immune attack; staphylokinase (SAK), which activates plasminogen to dissolve fibrin clots and promote bacterial dissemination; and nuclease (DNase), which degrades neutrophil extracellular traps (NETs) to facilitate immune evasion.
Major Diseases
S. aureus diseases fall into three broad categories. First, skin and soft tissue infections (SSTIs) are the most common and include folliculitis, furuncles (boils), carbuncles, impetigo, cellulitis, and abscesses. The emergence of CA-MRSA has led to increased severe skin infections in otherwise healthy young adults, frequently involving PVL-positive strains.
Second, invasive infections include bacteremia and sepsis, infective endocarditis, osteomyelitis, septic arthritis, pneumonia, brain abscess, and epidural abscess. SAB has a case-fatality rate of 15 to 30% and causes an estimated 300,000 deaths per year worldwide (JAMA, 2025). MRSA endocarditis carries a mortality rate of 30 to 37%.
Third, toxin-mediated diseases include staphylococcal food poisoning (preformed enterotoxins in contaminated food; onset 30 minutes to 8 hours; nausea, vomiting, abdominal cramps, diarrhea), SSSS (predominantly in neonates and young children), and toxic shock syndrome (TSS; sudden high fever, diffuse macular rash, hypotension, multi-organ involvement).
Biofilm Formation
S. aureus is highly proficient at forming biofilms on medical device surfaces. Biofilm development proceeds through initial attachment (mediated by surface proteins), accumulation (involving polysaccharide intercellular adhesin PIA/PNAG, extracellular DNA, and proteins), maturation, and dispersal. Bacteria within the biofilm matrix exhibit dramatically increased antibiotic tolerance, with minimum inhibitory concentrations (MICs) 10 to 1,000-fold higher than planktonic cells, and are shielded from immune cell penetration. Biofilm-associated infections occur on prosthetic joints, central venous catheters, prosthetic heart valves, vascular grafts, and orthopedic fixation devices, and typically require complete removal of the infected device for successful treatment.
6. Epidemiology and Transmission
Human Colonization
The anterior nares represent the primary ecological niche for S. aureus colonization. Additional colonization sites include the pharynx, perineum, axillae, groin, and skin surfaces (hands, chest, abdomen). According to the CDC (2024), approximately 20 to 30% of the population are persistent carriers, approximately 30% are intermittent carriers, and the remaining approximately 50% are non-carriers. MRSA nasal carriage prevalence is estimated at approximately 2% in the general population (CDC, 2025). Nasal carriage has been established as an independent risk factor for subsequent invasive infections including surgical site infections, bacteremia, and peritoneal dialysis-related infections.
Transmission Routes and Epidemiology
S. aureus is transmitted through direct contact (skin-to-skin), indirect contact (contaminated fomites and surfaces), and respiratory droplets. In healthcare settings, transmission occurs primarily via the hands of healthcare workers, contaminated medical equipment, and bed linens, making it one of the top five causes of nosocomial infections. In the community, CA-MRSA transmission is facilitated in settings involving close skin contact such as athletic facilities, correctional facilities, military barracks, and childcare centers.
Global MRSA prevalence varies substantially by region. A 2023 meta-analysis estimated the global pooled MRSA prevalence at approximately 14.7% (95% CI: 12.4-17.2%) of all S. aureus isolates. The GBD 2021 analysis revealed that MRSA-associated mortality is increasing worldwide: deaths directly attributable to MRSA rose from approximately 57,200 in 1990 to 130,000 in 2021, while deaths associated with MRSA rose from 261,000 to 550,000 over the same period (GBD 2021 AMR Collaborators, The Lancet, 2024). The United States reported approximately 120,000 SAB cases and 20,000 associated deaths annually (CDC, 2019).
Animal Hosts
S. aureus colonizes a wide range of animal species including companion animals (dogs, cats, horses), livestock (cattle, swine, poultry), and wildlife (hedgehogs). In dairy cattle, it is a leading cause of bovine mastitis. Livestock-associated MRSA (LA-MRSA, predominantly CC398) has emerged as a significant occupational hazard for farmers, veterinarians, and slaughterhouse workers. Bidirectional transmission between humans and companion animals has been well documented.
7. Immunity, Treatment, and Prevention
Immune Response
Host defense against S. aureus is primarily mediated by innate immunity. Physical barriers of the skin and mucous membranes, antimicrobial peptides (defensins, cathelicidins), and the complement system constitute the first line of defense. Neutrophils are the most critical innate immune effector cells, employing phagocytosis, the oxidative burst, and NET formation to kill bacteria. Adaptive immunity involves both antibody-mediated responses (opsonization, toxin neutralization) and T-cell responses (particularly Th17 cells), though S. aureus employs Protein A, capsular polysaccharides, complement inhibitory proteins, leukocidins, and superantigens to subvert immune defenses.
Antibiotic Treatment
MSSA infections are treated with anti-staphylococcal penicillins (nafcillin, oxacillin) or first-generation cephalosporins (cefazolin) for serious infections, and oral agents such as cephalexin or doxycycline for mild skin infections.
Vancomycin remains the standard first-line agent for serious MRSA infections. Alternative agents include daptomycin, linezolid, teicoplanin, trimethoprim-sulfamethoxazole (TMP-SMX), clindamycin, and ceftaroline (a fifth-generation cephalosporin with activity against MRSA). A 2024 network meta-analysis (Ju et al., Int J Infect Dis) found daptomycin to be the most effective agent for MRSA infections overall. For MRSA pneumonia, linezolid may be preferred due to superior lung tissue penetration. In August 2025, the DOTS trial published in JAMA demonstrated that dalbavancin, a long-acting lipoglycopeptide requiring only two intravenous doses, was noninferior to standard 4-to-6-week intravenous therapy for complicated S. aureus bacteremia, representing a potentially practice-changing advance in SAB management (NIH, 2025).
For VISA/VRSA infections, agents such as daptomycin, linezolid, quinupristin-dalfopristin, and ceftaroline are utilized, and infectious disease specialist consultation is strongly recommended.
Decolonization
Decolonization protocols are employed preoperatively and in high-risk carriers to prevent infection. The standard regimen consists of mupirocin 2% nasal ointment applied twice daily for 5 days combined with chlorhexidine 4% body wash. This approach has been demonstrated to significantly reduce surgical site infections.
Vaccine Development
No approved vaccine against S. aureus currently exists, despite decades of research. Multiple candidates, including StaphVAX (Nabi Biopharmaceuticals), V710 (Merck), and SA4Ag (Pfizer), have failed in Phase III clinical trials. Major obstacles to vaccine development include the organism's extensive immune evasion mechanisms, the presence of non-protective immune imprinting from prior natural exposure in most adults, and the requirement to stimulate both robust humoral and cellular immunity.
In December 2024, researchers at UC San Diego identified a mechanism by which S. aureus activates an immune-suppressive protein that halts the immune response, potentially explaining repeated vaccine failures. Their work showed that blocking this protein reactivated immune cell function and restored vaccine effectiveness in an animal model (UC San Diego, 2024). In the same month, LimmaTech Biologics' multivalent toxoid vaccine LBT-SA7 received FDA Fast Track designation for the prevention of skin and soft tissue infections, and the first participants in a Phase 1 clinical trial (NCT06719219) were vaccinated in February 2025 (CIDRAP, 2024; Contagion Live, 2025). In September 2025, a multivalent mRNA-LNP cocktail vaccine targeting five key virulence factors demonstrated superior efficacy in murine models of S. aureus infection, as reported in npj Vaccines.
8. Ecology and Environmental Role
Environmental Survival
S. aureus exhibits remarkable environmental resilience. It can survive NaCl concentrations up to 15%, enabling growth in salted and cured foods. It can persist on dry surfaces for weeks to months, contributing to persistent contamination in hospital environments. While optimal growth occurs at 30 to 37°C, the organism can survive across a range of 6 to 48°C. Under optimal aerobic conditions, the doubling time is approximately 20 to 35 minutes; under anaerobic conditions, it increases to approximately 80 minutes. In the human nose, the estimated doubling time is 1 to 3 hours (Krismer et al., 2019).
Microbial Community Interactions
Within the nasal niche, S. aureus engages in competitive and cooperative interactions with other members of the nasal microbiota, including Corynebacterium spp., Dolosigranulum pigrum, and S. epidermidis. Corynebacterium species produce free fatty acids through lipid degradation that inhibit S. aureus colonization. S. lugdunensis produces lugdunin, a novel thiazolidine-containing cyclic peptide antibiotic with potent activity against S. aureus, and has attracted attention as a potential probiotic-based decolonization strategy.
Food Safety
Staphylococcal food poisoning is an intoxication-type foodborne illness caused by ingestion of preformed enterotoxins in contaminated food, rather than by infection with live bacteria. The enterotoxins are remarkably heat-stable, resisting 100°C for 30 minutes, meaning that cooking cannot inactivate them once they have been produced. Foods commonly implicated include those handled extensively by humans, such as sandwiches, cream-based products, dairy products, and meat products. Inadequate refrigeration (above 5°C) allowing bacterial growth is the key risk factor.
9. Research History and Scientific Significance
Discovery and Early Research
The discovery of S. aureus is intertwined with the birth of modern bacteriology. In 1880, Alexander Ogston first observed grape-like clusters of bacteria in pus from surgical abscesses at Aberdeen Royal Infirmary in Scotland. Using mouse models, he demonstrated that these organisms were the causative agents of suppuration, contributing fundamentally to the germ theory of disease alongside Koch's postulates. In 1884, Rosenbach's formal description and naming based on colony pigmentation established species-level taxonomy. Over the following decades, key virulence factors including coagulase, hemolysins, Protein A, and enterotoxins were progressively identified.
A Model for Antibiotic Resistance Evolution
S. aureus serves as the paradigmatic model for understanding antibiotic resistance evolution. Penicillinase-producing resistant strains emerged within years of penicillin's introduction in the 1940s. Methicillin was introduced in 1959, and the first MRSA was reported in the United Kingdom in 1961. The first VRSA was isolated in Michigan, United States, in 2002. This historical sequence vividly illustrates the ongoing evolutionary arms race between antimicrobial agents and bacterial adaptation.
Current Research Directions
Current S. aureus research encompasses several major areas. In vaccine development, multivalent antigen strategies, mRNA-based platforms, and approaches to overcome immune suppression mechanisms are under active investigation. Alternative therapies under exploration include bacteriophage therapy, monoclonal antibody-based treatments (including multivalent nanobodies), anti-virulence strategies, and anti-biofilm agents. Epidemiologically, the blurring boundaries between HA-MRSA and CA-MRSA, the zoonotic potential of LA-MRSA, and global trends in antimicrobial resistance are key surveillance priorities. The 2025 DOTS trial demonstrating dalbavancin's noninferiority for complicated SAB represents a potential paradigm shift in treatment duration and patient management.
10. Comparison with Related Species
| Feature | S. aureus | S. epidermidis | S. saprophyticus | S. lugdunensis |
|---|---|---|---|---|
| Coagulase | Positive | Negative | Negative | Negative |
| Hemolysis | Beta-hemolytic | Non-hemolytic/gamma | Non-hemolytic/gamma | Beta-hemolytic |
| Colony color | Golden-yellow | White/gray | White/yellowish | Cream |
| Primary infections | Skin, pneumonia, sepsis, endocarditis | Device-related infections | Urinary tract infections | Endocarditis, skin infections |
| Pathogenicity | High | Low (opportunistic) | Low (urinary-specific) | Moderate to high |
| Biofilm formation | High | Very high | Low | Moderate |
| Primary niche | Anterior nares, skin | Skin (widespread) | Urogenital tract | Perineum |
| DNase | Positive | Negative | Negative | Positive |
S. epidermidis is the most abundant staphylococcal species on human skin and is generally nonpathogenic but is a leading cause of medical device-associated infections (catheters, prosthetic valves) due to its exceptional biofilm-forming capacity. S. saprophyticus is the second most common cause of urinary tract infections in young women. S. lugdunensis is notable for being a coagulase-negative species that can cause infections rivaling S. aureus in severity, particularly native valve endocarditis, and uniquely produces the antibiotic lugdunin that inhibits S. aureus colonization.
Fun Facts
The origin of methicillin resistance predates human antibiotic use by decades. A 2022 Nature study revealed that the mecC resistance gene evolved naturally on the skin of European hedgehogs coinfected with S. aureus and the fungus Trichophyton erinacei, which produces a natural beta-lactam antibiotic, as far back as the 1940s.
The golden pigment staphyloxanthin is not merely decorative; it acts as a molecular shield that neutralizes reactive oxygen species generated by immune cells, directly helping S. aureus survive inside the very white blood cells trying to kill it.
Under optimal laboratory conditions, S. aureus can divide every 20 minutes. Theoretically, a single cell could produce approximately 10 to the power of 21 cells in just 24 hours, though nutrient depletion and waste accumulation prevent this from happening in practice.
Staphylococcal enterotoxins can withstand boiling at 100°C for 30 minutes without being destroyed. Once these toxins have been produced in contaminated food, no amount of cooking can make it safe, which is why prevention must focus on stopping bacterial growth in the first place.
S. aureus can survive in salt concentrations up to 15% and persist on dry surfaces for weeks to months. This extraordinary environmental resilience makes it a persistent contaminant in hospitals and a concern in salt-preserved foods.
Protein A, a surface protein unique to S. aureus, binds antibodies 'backwards' by their Fc region, effectively coating the bacterium in a camouflage of wrongly oriented antibodies that cannot trigger immune attack, neutralizing a key defense mechanism.
According to the GBD 2021 analysis, MRSA-attributable deaths doubled from approximately 57,200 in 1990 to 130,000 in 2021, making MRSA the single deadliest pathogen-drug combination in the global antimicrobial resistance crisis.
Despite decades of intensive research and multiple Phase III clinical trials, no effective vaccine against S. aureus has ever been approved. In 2024, researchers discovered that S. aureus activates an immune-suppressive protein that directly sabotages vaccine-induced immunity, explaining years of failure.
Coagulase, the enzyme that gives S. aureus its 'coagulase-positive' distinction from other staphylococci, works by hijacking the host's own clotting system to encase the bacterium in a fibrin cloak, shielding it from immune cells like a biological suit of armor.
S. aureus can jump between humans and animals in both directions. Livestock-associated MRSA (LA-MRSA, CC398) arose in farm animals and spread to farmers, while pet dogs and cats commonly acquire MRSA from their owners and can serve as reinfection reservoirs.
In December 2024, the first-ever S. aureus vaccine candidate, LBT-SA7, received FDA Fast Track designation, and in February 2025, the first human participants were vaccinated in a Phase 1 clinical trial, marking a new chapter in the quest for a staphylococcal vaccine.
In August 2025, the DOTS trial showed that just two doses of the long-acting antibiotic dalbavancin could replace 4 to 6 weeks of daily intravenous therapy for complicated S. aureus bloodstream infections, potentially revolutionizing how this deadly infection is treated.
FAQ
Being an S. aureus carrier means the bacterium is present on your body, typically in the nostrils or on the skin, without causing any symptoms or disease. According to the CDC (2024), about 20 to 30% of people are persistent carriers, roughly 30% are intermittent carriers, and the remaining approximately 50% are non-carriers. Carriage itself is not a disease, but it increases the risk of developing an infection if the skin barrier is broken (through wounds, surgery, or medical device insertion) or if the immune system is weakened. Carriers can also transmit the bacteria to others. Nasal carriage has been identified as an independent risk factor for subsequent invasive infections such as surgical site infections and bacteremia.
MRSA (methicillin-resistant Staphylococcus aureus) carries the mecA gene, which encodes an altered penicillin-binding protein (PBP2a) with very low affinity for beta-lactam antibiotics. This makes MRSA resistant to methicillin and most other beta-lactam antibiotics (penicillins and cephalosporins). MSSA (methicillin-susceptible S. aureus) lacks this gene and can be effectively treated with standard anti-staphylococcal penicillins such as oxacillin or first-generation cephalosporins like cefazolin. MRSA infections require different antibiotics such as vancomycin or daptomycin, are more expensive to treat, and are associated with higher mortality rates. For example, MRSA bacteremia mortality ranges from 20 to 50%, generally exceeding that of MSSA bacteremia.
Symptoms vary widely depending on the type and location of infection. Skin infections present as redness, swelling, warmth, pain, and pus-filled boils or abscesses. Food poisoning manifests as nausea, vomiting, abdominal cramps, and diarrhea, with a very rapid onset (30 minutes to 8 hours after eating contaminated food). Invasive infections can cause high fever and chills (bacteremia/sepsis), shortness of breath and productive cough (pneumonia), joint pain and swelling (septic arthritis), chest pain and heart murmur (endocarditis), and bone pain (osteomyelitis). Toxic shock syndrome presents with sudden high fever, a diffuse sunburn-like rash, dangerously low blood pressure, and multi-organ dysfunction. Bacteremia and sepsis can be life-threatening, with a case-fatality rate of 15 to 30%.
In the community, key prevention measures include washing hands frequently with soap and water or using alcohol-based hand sanitizer, keeping cuts and wounds clean, dry, and covered with a bandage until healed, avoiding sharing personal items such as towels, razors, and athletic clothing, and cleaning shared surfaces (gym equipment, etc.) before and after use. In healthcare settings, prevention strategies include strict hand hygiene compliance by healthcare workers, contact precautions (gowns and gloves) for patients with known MRSA, environmental surface disinfection, active surveillance screening for MRSA in high-risk patients, and preoperative decolonization with mupirocin nasal ointment and chlorhexidine body wash to reduce surgical site infections.
There is no approved vaccine against S. aureus as of 2025. Several candidates, including StaphVAX, V710, and SA4Ag, failed in Phase III clinical trials. However, promising developments are underway. In December 2024, LimmaTech Biologics' multivalent toxoid vaccine LBT-SA7 received FDA Fast Track designation, and the first participants were vaccinated in a Phase 1 trial in February 2025. Additionally, mRNA-based multivalent vaccines have shown encouraging results in preclinical models. A key challenge is that S. aureus employs multiple immune evasion mechanisms and most adults already have non-protective immune imprinting from prior exposure, which the bacterium exploits to suppress vaccine-induced immunity. Researchers at UC San Diego identified a mechanism for this suppression in 2024, offering a potential path to overcome it.
S. aureus food poisoning is an intoxication rather than an infection. The bacteria produce enterotoxins while growing in food before it is consumed, so the toxin is already preformed at the time of ingestion. Because the body does not need to wait for bacterial multiplication, symptoms begin very rapidly, typically within 30 minutes to 8 hours. Furthermore, staphylococcal enterotoxins are remarkably heat-stable, withstanding 100°C (212°F) for 30 minutes, meaning that cooking cannot destroy them once produced. This is why prevention focuses on proper food handling, hand hygiene, and keeping food refrigerated at or below 5°C (41°F) to prevent bacterial growth and toxin production in the first place.
S. aureus excels at forming biofilms on implanted medical device surfaces. When a device is inserted into the body, it quickly becomes coated with host proteins (fibrinogen, fibronectin), to which S. aureus adheres through its MSCRAMMs surface proteins. The bacteria then produce an extracellular matrix composed of polysaccharide intercellular adhesin (PIA/PNAG), extracellular DNA, and proteins, forming a mature biofilm. Within this structure, bacteria become metabolically dormant and are shielded from both antibiotics (MICs increase 10 to 1,000-fold) and immune cells. This is why biofilm-associated device infections, affecting prosthetic joints, catheters, heart valves, and orthopedic hardware, are extremely difficult to treat with antibiotics alone and typically require surgical removal of the infected device.
The golden-yellow color comes from staphyloxanthin, a carotenoid pigment produced by the bacterium and incorporated into its cell membrane. Staphyloxanthin is far more than a cosmetic feature; it functions as a critical virulence factor. The pigment acts as an antioxidant that scavenges reactive oxygen species (ROS) and singlet oxygen generated by host neutrophils during the oxidative burst, the primary mechanism by which these immune cells kill engulfed bacteria. Mutant strains engineered to lack staphyloxanthin production are significantly more susceptible to neutrophil killing, confirming that the golden pigment directly contributes to immune evasion and bacterial survival within the host.
The rise of MRSA is driven by multiple factors. The overuse and misuse of antibiotics in both human medicine and agriculture provide selective pressure favoring resistant strains. In healthcare settings, inadequate infection control practices facilitate cross-transmission via healthcare workers' hands, contaminated medical equipment, and environmental surfaces. The emergence of CA-MRSA in the 1990s expanded the problem beyond hospitals into the general community, affecting otherwise healthy individuals. Antibiotic use in livestock farming has driven the expansion of LA-MRSA (primarily CC398), which can transmit to humans. According to the GBD 2021 analysis, deaths attributable to MRSA more than doubled between 1990 and 2021, rising from 57,200 to 130,000 globally, underscoring the accelerating threat.
Yes, S. aureus, including MRSA, can be transmitted bidirectionally between humans and companion animals such as dogs, cats, and horses. Pets can acquire S. aureus from their owners and then serve as a reservoir that re-transmits the bacteria back to humans or other household members. Livestock-associated MRSA (LA-MRSA, predominantly CC398) is particularly prevalent in farm animals (especially swine) and represents a significant occupational risk for farmers, veterinarians, and slaughterhouse workers. This human-animal interface is an important aspect of MRSA epidemiology and underscores the need for a One Health approach to surveillance and infection control.
In August 2025, the landmark DOTS trial published in JAMA demonstrated that dalbavancin, a long-acting lipoglycopeptide antibiotic requiring only two intravenous doses, was noninferior to standard-of-care therapy (which typically requires 4 to 6 weeks of daily intravenous antibiotics) for complicated S. aureus bacteremia. This represents a potentially practice-changing development, as it could dramatically simplify treatment, reduce hospitalization duration, enable outpatient management, and improve patients' quality of life while maintaining equivalent clinical outcomes. However, further research is ongoing to define the optimal patient populations for this approach.
📚References
- Rosenbach FJ. Mikroorganismen bei den Wund-Infections-Krankheiten des Menschen. J.F. Bergmann, Wiesbaden, 1884.
- Ogston A. Report upon micro-organisms in surgical diseases. British Medical Journal. 1881;1(1054):369-375. https://doi.org/10.1136/bmj.1.1054.369.b
- Tong SY, Davis JS, Eichenberger E, Holland TL, Fowler VG Jr. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clinical Microbiology Reviews. 2015;28(3):603-661. https://doi.org/10.1128/CMR.00134-14
- Lowy FD. Staphylococcus aureus infections. New England Journal of Medicine. 1998;339(8):520-532. https://doi.org/10.1056/NEJM199808203390806
- GBD 2019 Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The Lancet. 2022;399(10325):629-655. https://doi.org/10.1016/S0140-6736(21)02724-0
- GBD 2021 Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance 1990-2021: a systematic analysis with forecasts to 2050. The Lancet. 2024;404(10459):1199-1226. https://doi.org/10.1016/S0140-6736(24)01867-1
- CDC. Vital Signs: Epidemiology and Recent Trends in Methicillin-Resistant Staphylococcus aureus Bloodstream Infections. MMWR Morb Mortal Wkly Rep. 2019;68(9):214-219. https://www.cdc.gov/mmwr/volumes/68/wr/mm6809e1.htm
- CDC. Staphylococcus aureus Basics. Centers for Disease Control and Prevention. 2024. https://www.cdc.gov/staphylococcus-aureus/about/index.html
- Eiamphungporn W, Schauss AC, Heine D, et al. Emergence of methicillin resistance predates the clinical use of antibiotics. Nature. 2022;602:135-141. https://doi.org/10.1038/s41586-021-04265-w
- Foster TJ. Antibiotic resistance in Staphylococcus aureus. Current status and future prospects. FEMS Microbiology Reviews. 2017;41(3):430-449. https://doi.org/10.1093/femsre/fux007
- Wertheim HF, Melles DC, Vos MC, et al. The role of nasal carriage in Staphylococcus aureus infections. The Lancet Infectious Diseases. 2005;5(12):751-762. https://doi.org/10.1016/S1473-3099(05)70295-4
- Otto M. MRSA virulence and spread. Cellular Microbiology. 2012;14(10):1513-1521. https://doi.org/10.1111/j.1462-5822.2012.01832.x
- Krismer B, Abt C, Taxis K, et al. Rapid cell division of Staphylococcus aureus during colonization of the human nose. mBio. 2019;10(2):e00285-19. https://doi.org/10.1128/mBio.00285-19
- Liu C, Bayer A, Cosgrove SE, et al. Clinical Practice Guidelines by the Infectious Diseases Society of America for the Treatment of Methicillin-Resistant Staphylococcus aureus Infections in Adults and Children. Clinical Infectious Diseases. 2011;52(3):e18-e55. https://doi.org/10.1093/cid/ciq146
- Ju G, Han Y, Chen J, et al. Comparative effectiveness and safety of six antibiotics in treating MRSA infections: a network meta-analysis. International Journal of Infectious Diseases. 2024;145:107076. https://doi.org/10.1016/j.ijid.2024.107076
- WHO. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics. World Health Organization. 2017.
- Skerman VBD, McGowan V, Sneath PHA. Approved lists of bacterial names. International Journal of Systematic Bacteriology. 1980;30:225-420.
- LimmaTech Biologics. FDA Fast Track Designation for LBT-SA7. Press Release. December 19, 2024. https://lmtbio.com
- CDC/NIH. Biosafety in Microbiological and Biomedical Laboratories (BMBL), 6th Edition. 2020. https://www.cdc.gov/labs/pdf/SF__19_308133-A_BMBL6_00-BOOK-WEB-final-3.pdf
- Holland TL, Cosgrove SE, Doernberg SB, et al. Dalbavancin for Treatment of Staphylococcus aureus Bacteremia. JAMA. 2025. https://doi.org/10.1001/jama.2025.12437
Gallery
4 images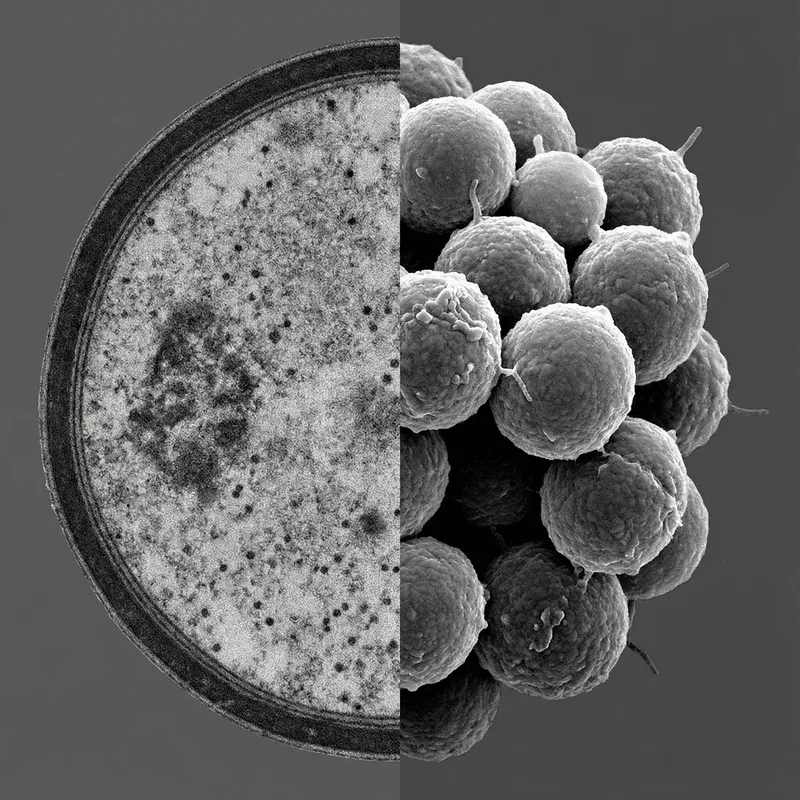 Staphylococcus aureus
Staphylococcus aureusStaphylococcus aureus
 Staphylococcus aureus
Staphylococcus aureusStaphylococcus aureus
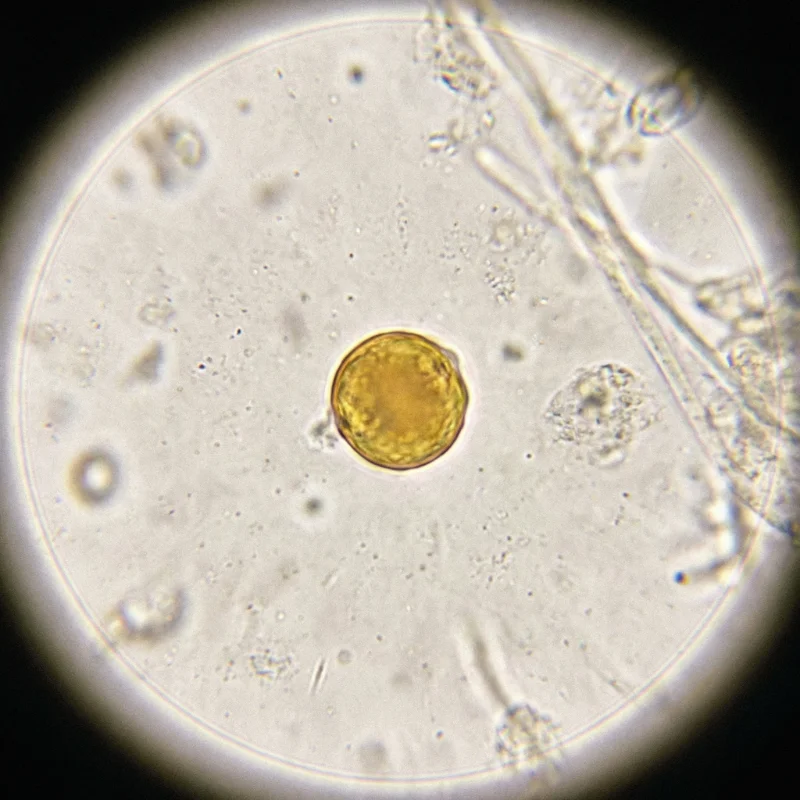 Staphylococcus aureus
Staphylococcus aureusStaphylococcus aureus
 Staphylococcus aureus
Staphylococcus aureusStaphylococcus aureus