Slipper Animalcule
Protists · Amoebozoa
Paramecium caudatum
Name meaning: "The genus name Paramecium derives from the Greek word paramēkēs meaning 'oblong' or 'oval,' referring to the elongated cell shape. The species epithet caudatum comes from the Latin word for 'tailed,' describing the pointed posterior end of the cell."
Discovery
Microorganism traits
Classification
Habitat
Classification history
First observed by Leeuwenhoek around 1674 and clearly described by Huygens in 1678. The term Paramecium was first used by John Hill in 1752. Otto Friedrich Müller formally introduced the genus into Linnaean taxonomy in 1773. The species P. caudatum was first described by Christian Gottfried Ehrenberg in 1833. Modern molecular phylogenetics places it within SAR supergroup > Alveolata > Ciliophora > Oligohymenophorea > Peniculida > Parameciidae (NCBI Taxon ID 5885).
Clinical significance
Non-pathogenic free-living protist with no association with human disease. Significant as a model organism in biology research and as an indicator species in ecotoxicology.
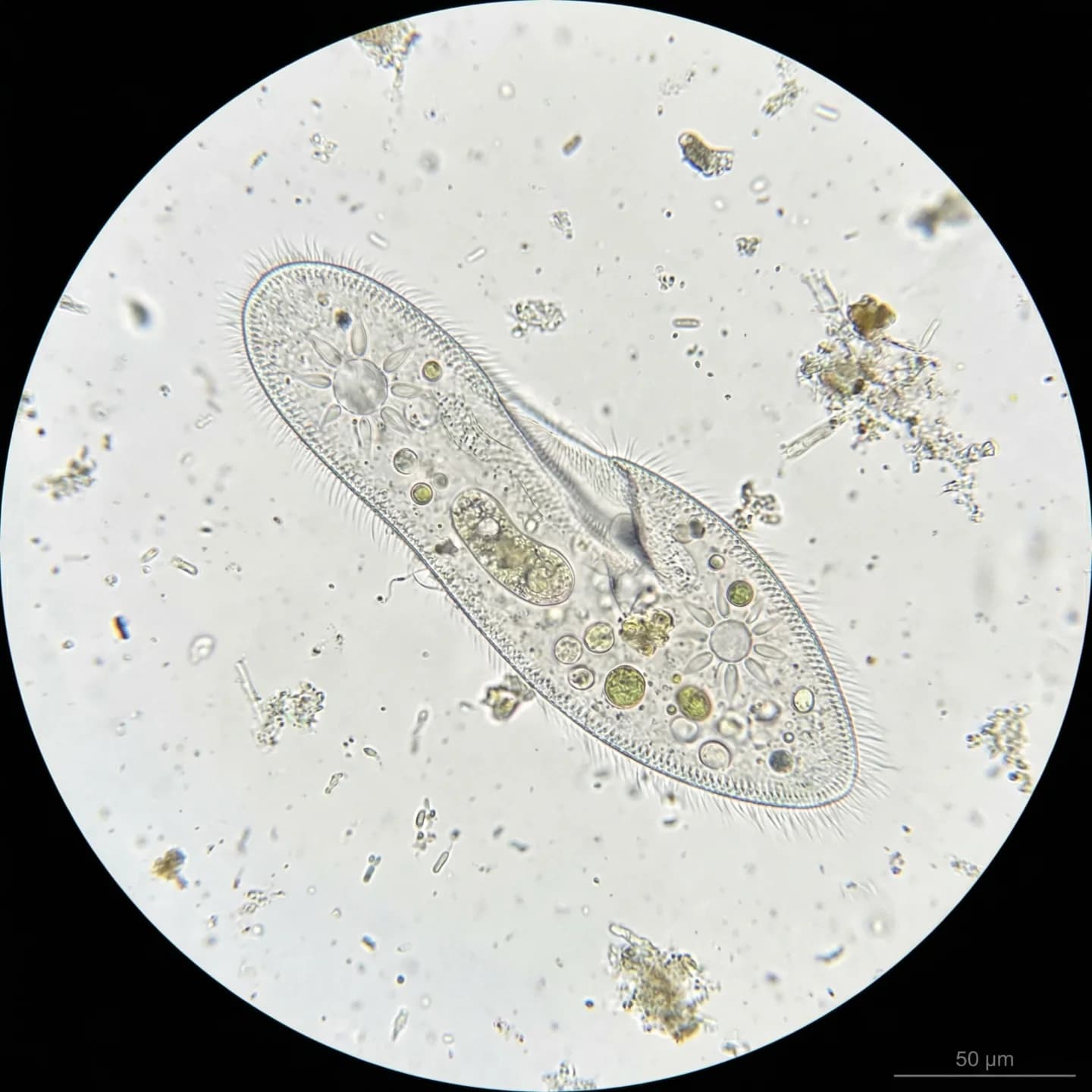

Paramecium caudatum Ehrenberg, 1833 is a unicellular eukaryote belonging to the phylum Ciliophora, class Oligohymenophorea, order Peniculida, and family Parameciidae. It is a free-living, non-pathogenic protist with a cosmopolitan distribution across freshwater, brackish, and occasionally marine environments worldwide. Cells measure 170–330 μm in length (typically 200–300 μm) and are covered by approximately 4,000 motile cilia arranged in longitudinal rows, which beat in coordinated metachronal waves to propel the organism in a spiraling path through the water (Bouhouche et al., 2022). The species epithet "caudatum" is Latin for "tailed," referring to the distinctly pointed posterior end, while the genus name derives from the Greek "paramēkēs" (oblong). Early microscopists likened the overall cell shape to a slipper, giving rise to the enduring nickname "slipper animalcule."
P. caudatum is one of the most important model organisms in biology, often called the "white rat of the Ciliophora" (Van Houten, 2023). It was the key experimental subject in Georgy Gause's landmark 1934 experiments that established the competitive exclusion principle, a foundational concept in ecology (Gause, 1934). The species has also been central to research on nuclear dimorphism, programmed genome rearrangement, ciliary motility, and ion channel-mediated behavior. Notably, its calcium-based action potentials that trigger complex avoidance reactions have earned it the distinctive label of "swimming neuron" (Brette, 2021).
From a biosafety perspective, P. caudatum is a BSL-1 non-pathogenic organism that can be cultured and handled without special containment requirements. It is not listed on the IUCN Red List and is globally abundant, with no current conservation concerns.
1. Overview
The genus name Paramecium derives from the Greek word "paramēkēs," meaning oblong or oval, while the species epithet "caudatum" comes from the Latin for "tailed," reflecting the tapered posterior end of the cell. The colloquial English name "slipper animalcule" originated with the French microscopist Louis Joblot, who called this creature "Chausson" (slipper), and the term persisted through the 18th and 19th centuries as a popular designation for Paramecium species (Merriam-Webster).
P. caudatum is currently registered in NCBI Taxonomy under Taxon ID 5885 and is classified within the SAR supergroup under Alveolata. More than 15 morphospecies are recognized within the genus Paramecium (Fokin, 2010). The most distinctive feature of this species, shared with all ciliates, is nuclear dimorphism: each cell contains a polyploid macronucleus responsible for somatic functions and a single diploid micronucleus that serves as the germline repository.
2. Taxonomy and Phylogeny
The modern classification of P. caudatum is built on molecular phylogenetic evidence and places it within the SAR supergroup (Stramenopila–Alveolata–Rhizaria). Within the Alveolata, the ciliates (Ciliophora) form a sister group to the dinoflagellates (Dinoflagellata) and the apicomplexans (Apicomplexa). The full taxonomic hierarchy is as follows: Domain Eukaryota, Clade SAR, Superphylum Alveolata, Phylum Ciliophora, Class Oligohymenophorea, Order Peniculida, Family Parameciidae, Genus Paramecium Müller, 1773, Species P. caudatum Ehrenberg, 1833.
Phylogenetic studies of the genus Paramecium have relied primarily on 18S rRNA gene sequences, ITS regions, and mitochondrial COI gene sequences. Strüder-Kypke et al. (2000) demonstrated through 18S rRNA-based phylogenetic analysis that Paramecium species form at least four clades, with the P. aurelia subgroup occupying the most derived position. P. caudatum is sister to the P. aurelia species complex, and the two lineages share only the most ancient of three whole-genome duplication (WGD) events (McGrath et al., 2014).
The genus Paramecium was formally introduced into Linnaean nomenclature in 1773 by the Danish zoologist Otto Friedrich Müller, although the term itself was first used by John Hill in 1752. The species P. caudatum was first described in 1833 by the German naturalist Christian Gottfried Ehrenberg (Fokin, 2010). The very first observations of Paramecium are attributed to Antonie van Leeuwenhoek around 1674, and a clear description was provided in a 1678 letter from Christiaan Huygens to the Royal Society.
Within the genus, P. caudatum is a relatively large species distinguished by its single micronucleus, in contrast to the two micronuclei found in the P. aurelia species complex. Other notable congeners include P. aurelia (a complex of 15 cryptic species), P. bursaria (harboring green algal endosymbionts), P. multimicronucleatum (possessing 3–7 micronuclei), and P. tetraurelia (the primary genomic model for the genus).
3. Morphology and Structure
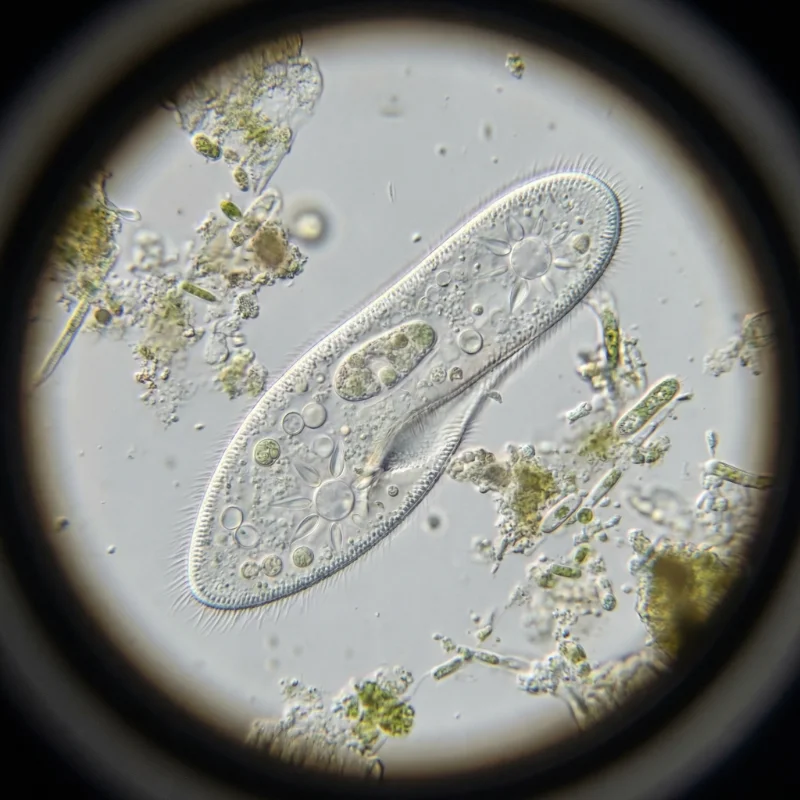
P. caudatum is a spindle-shaped (fusiform) unicellular organism, with a rounded anterior end and a characteristically pointed posterior end. Cell length ranges from 170 to 330 μm, typically 200–300 μm, placing it among the larger species in the genus (Fokin, 2010). Cell width is approximately 45–80 μm (Wichterman, 2012). The relatively large size means that individual cells can be faintly visible to the naked eye as tiny white specks and are easily observed under low-magnification light microscopy.
The cell surface is enveloped by the pellicle, composed of the plasma membrane underlain by alveolar sacs. Beneath the pellicle, thousands of spindle-shaped extrusomes called trichocysts are densely arrayed and can be explosively discharged as a defense mechanism against predators (Harumoto & Miyake, 1991). On the ventral side of the anterior half, the oral groove (vestibulum) leads into the deep buccal cavity, which terminates at the cytostome and cytopharynx, forming the feeding apparatus.
The entire cell surface is uniformly covered by approximately 4,000 motile cilia arranged in longitudinal rows (Bouhouche et al., 2022). Individual cilia measure 10–12 μm in length and approximately 0.27 μm in diameter, each possessing a 9+2 microtubule axoneme originating from a basal body. At the posterior end, caudal cilia measuring 10–20 μm in length extend outward with a characteristic curvature. P. caudatum possesses two contractile vacuoles, one anterior and one posterior (Patterson, 1980), each connected to radially arranged collecting canals that give them a distinctive star-shaped appearance. These organelles regulate osmotic pressure by expelling excess water that continuously enters the cell in hypotonic freshwater environments.
The most distinctive structural feature of P. caudatum is nuclear dimorphism (Mikami, 1988). The macronucleus is a large, kidney-shaped polyploid nucleus that regulates everyday somatic functions including metabolism, nutrition, and respiration, with extremely high transcriptional activity. It divides by amitosis during binary fission. The micronucleus is a small, compact diploid nucleus that undergoes meiosis and mitosis during sexual reproduction (conjugation), preserving and transmitting germline genetic information. P. caudatum possesses a single micronucleus, a key morphological distinction from P. aurelia, which has two.
4. Genome and Molecular Biology
Like all ciliates, P. caudatum maintains two distinct nuclear genomes. The germline genome (micronucleus) is estimated at approximately 1,300–5,500 Mb, representing one of the largest germline genomes among ciliates (McGrath et al., 2014; Furrer et al., 2024). The somatic genome (macronucleus) retains only about 2% of the germline genome through programmed DNA elimination, one of the most extreme genome rearrangement events known in biology.
The macronuclear genome assembly of P. caudatum, published by McGrath et al. (2014), comprises approximately 30.5 Mb distributed across 1,202 scaffolds, with 18,509 annotated protein-coding genes. This is less than half the genome size of P. aurelia species complex members (68–77 Mb) and approximately half their gene count (~34,000–40,000 genes). The genomic GC content is 28.2%, and the average intergenic region length is only 110 bp, making it one of the most compact eukaryotic genomes known. These data are available through ParameciumDB.
Phylogenomic analysis has confirmed that P. caudatum shares only the most ancient of three WGD events with the P. aurelia lineage, estimated to have occurred approximately 1.5 billion years ago based on synonymous substitution rates (McGrath et al., 2014). The two more recent WGDs occurred exclusively in the P. aurelia lineage after its divergence from P. caudatum. Remarkably, P. caudatum retains about 16% of duplicate genes from the ancient WGD, roughly twice the retention rate observed in P. aurelia species (~8%), supporting the hypothesis that subsequent WGDs relax dosage constraints on ancient duplicates. A 2025 study analyzing macronuclear genomes of five additional Paramecium species revealed unexpected diversity in genome sizes ranging from 47.78 to 113.16 Mb (Li et al., 2025).
The mitochondrial genome is linear and was fully sequenced by Barth & Behnke (2011), revealing significant shifts in nucleotide composition and codon usage compared to P. tetraurelia, indicative of active mitochondrial genome evolution within the genus.
5. Ecology and Environmental Role
P. caudatum exhibits a cosmopolitan distribution, with isolates identified on every continent except Antarctica (Fokin, 2010). Its primary habitats are freshwater environments including ponds, lakes, streams, reservoirs, and irrigation channels, with particularly high population densities in stagnant waters rich in decaying organic matter. It has also been found in brackish and marine environments, though at lower frequencies. The organism commonly inhabits the mud-water interface of littoral freshwater environments.
As a heterotroph, P. caudatum feeds on bacteria, yeasts, unicellular algae, and fine organic particles. Cilia in the oral groove beat coordinately to draw food particles and water into the cell, where food vacuoles form at the cytopharynx. Intracellular digestion occurs through fusion with lysosomes during cytoplasmic streaming (cyclosis), and undigested residues are expelled through the cytoproct. This bacterivorous activity plays an important role in freshwater microbial food webs, contributing to the regulation of bacterial community structure and nutrient recycling.
Optimal growth temperature for P. caudatum is approximately 24–29°C, with a thermal range for growth spanning 7–35.5°C (Krenek et al., 2012). Optimal pH is in the range of 6.0–7.0, and under hypoxic conditions, pH 4.7–6.7 has been reported as most favorable for survival (Heydarnejad, 2008). As an aerobic organism, it requires dissolved oxygen but can tolerate brief periods of hypoxia. In freshwater environments, osmotic balance is maintained through the two contractile vacuoles, and the organism responds to environmental gradients through chemotaxis and thermotaxis.
The primary predator of P. caudatum is the carnivorous ciliate Didinium nasutum. The Didinium–Paramecium system is a classical model for predator–prey interaction studies and has been widely used to experimentally test Lotka–Volterra population dynamics models. P. caudatum defends itself by explosive discharge of trichocysts, which can propel the cell away from a predator's attack range and neutralize toxins (Harumoto & Miyake, 1991; Miyake, 1996).
P. caudatum is also known to host several species of intracellular endosymbiotic bacteria. The most prominent are members of the genus Holospora (Alphaproteobacteria): H. obtusa is macronucleus-specific, while H. undulata and H. recta are micronucleus-specific symbionts (Görtz et al., 2009). These endosymbionts enter the host through food vacuoles and migrate to their target nucleus. Notably, Holospora infection has been reported to confer increased resistance to heat stress and oxidative stress on the host (Hori & Fujishima, 2003).
6. Research History and Scientific Significance
The earliest observations of Paramecium are attributed to Antonie van Leeuwenhoek around 1674, with a definitive description provided in a 1678 letter from Christiaan Huygens to the Royal Society. Louis Joblot named the organism "Chausson" (slipper), establishing the lasting "slipper animalcule" nickname. John Hill first used the term "Paramecium" in 1752, and Otto Friedrich Müller formally introduced it into Linnaean taxonomy in 1773. The species P. caudatum was first described by Christian Gottfried Ehrenberg in 1833 and has since become one of the most intensively studied protists in biological research.
The most celebrated milestone in P. caudatum research history is Georgy Gause's 1934 formulation of the competitive exclusion principle (Gause's law). By co-culturing P. caudatum and P. aurelia in the same medium, Gause demonstrated that P. aurelia, being the more efficient competitor for the same resources, consistently drove P. caudatum to local extinction. This experiment established the fundamental ecological principle that two species competing for the same limiting resource cannot coexist indefinitely, and remains a landmark in the history of ecology (Gause, 1934).
As a model organism, P. caudatum has contributed to multiple fields. In cell biology and genetics, it has been used to study nuclear dimorphism, programmed genome rearrangement, and non-Mendelian inheritance. In ciliary biology, its approximately 4,000 motile cilia provide an exceptional system for studying ciliary beat mechanics and basal body organization, with direct relevance to human ciliopathies (Valentine & Van Houten, 2021). In neurophysiology, its calcium-based action potentials that trigger sophisticated avoidance reactions have made it a unique single-celled model for studying ion channel-mediated behavioral regulation (Brette, 2021). In ecotoxicology, it serves as an indicator organism for bioassays assessing environmental pollutant toxicity. In education, its relatively large size, active motility, and ease of culture make it a standard teaching organism for microscopy and cell biology laboratories worldwide.
The complete sequencing of the P. caudatum macronuclear genome in 2014 (McGrath et al., 2014) opened new avenues for comparative genomics within the genus. In 2024, RNA interference (RNAi) and protein localization techniques were successfully applied to P. caudatum, significantly expanding the molecular toolkit available for this species (Furrer et al., 2024).
7. Reproduction and Life Cycle
The primary mode of reproduction in P. caudatum is transverse binary fission. Under optimal conditions (24–28°C), cells can divide 2–3 times per day, with each division cycle taking approximately 6–8 hours (Wichterman, 2012). During fission, the macronucleus divides by amitosis while the micronucleus undergoes mitosis. The cell then divides transversely, with each daughter cell receiving one macronucleus and one micronucleus.
Conjugation is the sexual process in P. caudatum, generating genetic diversity and preventing clonal aging. Two cells of compatible mating types temporarily fuse and exchange genetic material. During this process, the micronucleus of each cell undergoes meiosis to produce haploid gametic nuclei, one of which migrates to the partner cell. After nuclear fusion, new macronuclei and micronuclei differentiate. P. caudatum can also undergo cytogamy, a form of self-fertilization that can be experimentally induced (Yanagi & Haga, 1998).
The immaturity period of a P. caudatum clone is approximately 60 fissions, and if only asexual reproduction continues without conjugation or autogamy, clonal death occurs after approximately 600 fissions (Uezu et al., 2009). Total clonal lifespans have been reported to range from 250 to 750 fissions across different lineages (Komori et al., 1979). Nuclear reorganization through conjugation or self-fertilization produces a rejuvenation effect that extends clonal lifespan, making this system a valuable model for studying the evolutionary advantages of sexual reproduction.
8. Comparison with Related Species
| Species | Cell length (μm) | Micronuclei | Morphological features | WGD events | Notable characteristics |
|---|---|---|---|---|---|
| P. caudatum | 170–330 | 1 | Fusiform, pointed posterior | 1 | Gause's experiment subject, genome ~30.5 Mb |
| P. aurelia (complex) | 80–170 | 2 | Oval, rounded posterior | 3 | 15 cryptic species, genome 68–77 Mb |
| P. bursaria | 80–150 | 1 | Ovoid, wide oral groove | — | Chlorella endosymbionts, green coloration |
| P. multimicronucleatum | 200–350 | 3–7 | Fusiform, large | — | Multiple micronuclei |
| P. tetraurelia | 100–150 | 2 | Fusiform | 3 | Standard genomic model species |
In Gause's competitive exclusion experiments, P. aurelia consistently outcompeted P. caudatum due to its more efficient utilization of shared resources. By contrast, P. bursaria can coexist with other Paramecium species because its endosymbiotic Chlorella algae provide an additional carbon source through photosynthesis, effectively partitioning the niche. In terms of genome size, the macronuclear genome of P. caudatum (~30.5 Mb) is less than half that of P. aurelia species (68–77 Mb), a difference attributable to two additional WGDs that occurred exclusively in the P. aurelia lineage (McGrath et al., 2014).
Fun Facts
Despite being a single-celled organism, Paramecium caudatum coordinates approximately 4,000 cilia in metachronal waves to swim in elegant spirals. This remarkable motor control has led some scientists to call it a 'swimming neuron' (Brette, 2021).
The avoidance reaction of P. caudatum is triggered by calcium-based action potentials strikingly similar to the signaling mechanisms used by neurons in the human brain—complex electrical signaling in a cell with no brain at all.
In 1934, Russian ecologist Georgy Gause used P. caudatum and P. aurelia to formulate the competitive exclusion principle (Gause's law), establishing one of the foundational concepts of modern ecology (Gause, 1934).
The germline genome of P. caudatum can be as large as 5,500 Mb, yet the working somatic genome retains only about 2% (~30.5 Mb) after programmed DNA elimination. The discarded 98% represents one of the most extreme genome rearrangements in any organism (McGrath et al., 2014).
The macronuclear genome of P. caudatum packs 18,509 genes into just 30.5 Mb with an average intergenic distance of only 110 base pairs—one of the most compact eukaryotic genomes ever described (McGrath et al., 2014).
P. caudatum uses two star-shaped contractile vacuoles to pump out excess water that constantly floods into the cell in freshwater environments. Without this continuous pumping, the cell would swell and burst (Patterson, 1980).
Thousands of spindle-shaped trichocysts are loaded beneath the cell surface like microscopic torpedoes. When threatened, P. caudatum can discharge them explosively within milliseconds to escape predators or neutralize their toxins (Miyake, 1996).
The primary predator of P. caudatum, the carnivorous ciliate Didinium nasutum, can swallow a Paramecium whole despite being much smaller. This predator–prey pair is a classical model for testing Lotka–Volterra population dynamics.
P. caudatum can host intracellular Holospora bacteria. Surprisingly, infected cells become more resistant to heat stress and oxidative stress than uninfected ones—a case where parasitic infection may actually benefit the host (Hori & Fujishima, 2003).
A P. caudatum clone reproducing only asexually dies after about 600 fissions. But if cells undergo conjugation (sex), they experience 'rejuvenation' and their lifespan resets—a powerful demonstration of why sexual reproduction persists in evolution (Uezu et al., 2009).
The name 'slipper animalcule' originated with French microscopist Louis Joblot, who called Paramecium 'Chausson' (slipper). In Korean, the organism is called 짚신벌레 (jipshin-beolle), also named after a type of traditional straw slipper—different cultures independently chose the same footwear metaphor.
FAQ
Paramecium caudatum cells measure 170–330 μm (micrometers) in length, typically 200–300 μm. This is approximately 0.2–0.3 mm, large enough to be faintly visible to the naked eye as a tiny white speck. It is one of the larger species in the genus Paramecium, roughly twice the size of P. aurelia (80–170 μm) (Fokin, 2010).
P. caudatum moves using approximately 4,000 cilia that cover its entire cell surface (Bouhouche et al., 2022). These cilia beat in coordinated metachronal waves, propelling the organism in a spiraling path while simultaneously rotating along its long axis. The average swimming speed is approximately 1,000–2,000 μm/s (about 1–2 mm per second), covering 5–10 body lengths each second (Brette, 2021).
P. caudatum is a heterotroph that feeds primarily on bacteria, yeasts, and unicellular algae. Cilia lining the oral groove beat coordinately to sweep food particles and water into the cell through the cytostome. Food vacuoles form at the base of the cytopharynx, and intracellular digestion proceeds through fusion with lysosomes. Undigested material is expelled through the cytoproct.
The primary mode of reproduction is asexual binary fission. Under optimal conditions, cells can divide 2–3 times per day, with each cycle taking about 6–8 hours (Wichterman, 2012). P. caudatum also undergoes conjugation, a sexual process in which two cells of compatible mating types temporarily fuse and exchange genetic material. This generates genetic diversity and prevents clonal aging.
All ciliates, including P. caudatum, exhibit nuclear dimorphism. The macronucleus is a large polyploid nucleus that controls everyday somatic functions such as metabolism, nutrition, and respiration. The micronucleus is a small diploid nucleus that preserves germline genetic information and undergoes meiosis during sexual reproduction. This division of labor allows the cell to maintain high transcriptional activity while preserving an intact germline genome (Catania et al., 2024).
Clonal lifespan in P. caudatum is measured in number of fissions. Without conjugation or self-fertilization, a clone undergoes clonal death after approximately 600 fissions (Uezu et al., 2009). Nuclear reorganization through conjugation produces a rejuvenation effect that extends the clonal lifespan. Total lifespans ranging from 250 to 750 fissions have been reported across different lineages (Komori et al., 1979).
P. caudatum is found on every continent except Antarctica. It is most abundant in freshwater environments such as ponds, lakes, streams, reservoirs, and irrigation channels, particularly in stagnant waters rich in bacteria and decaying organic matter. It is occasionally found in brackish and marine environments as well (Fokin, 2010).
P. caudatum exhibits a characteristic 'avoidance reaction': when it encounters mechanical, chemical, or thermal stimuli, calcium ions rush into the cell, reversing the direction of ciliary beating and causing the organism to swim backward, reorient, and then resume forward swimming (Brette, 2021). Additionally, it can explosively discharge trichocysts—spindle-shaped projectiles stored beneath the pellicle—to escape predators or neutralize their toxins (Miyake, 1996).
P. caudatum is called the 'white rat of the Ciliophora' due to its extensive use as a model organism. It was the key subject of Gause's 1934 competitive exclusion experiments (Gause, 1934) and has been central to research on nuclear dimorphism, genome rearrangement, ciliary biology relevant to human ciliopathies (Valentine & Van Houten, 2021), and ion channel-mediated behavioral regulation (Brette, 2021). Its large size, active motility, and ease of culture also make it a staple of biology education.
The germline genome (micronucleus) of P. caudatum is enormous—estimated at 1,300–5,500 Mb—yet the somatic genome (macronucleus) retains only about 2% of it (~30.5 Mb) after programmed DNA elimination, one of the most extreme genome rearrangements known. The macronuclear genome contains 18,509 genes with an average intergenic distance of just 110 bp, making it one of the most compact eukaryotic genomes ever described (McGrath et al., 2014).
The genus Paramecium contains more than 15 recognized morphospecies (Fokin, 2010). P. aurelia is a complex of 15 cryptic species, smaller than P. caudatum (80–170 μm) with two micronuclei. P. bursaria harbors endosymbiotic green algae (Chlorella), giving it a green color. P. multimicronucleatum is similar in size but possesses 3–7 micronuclei, and P. tetraurelia is the standard genomic model for the genus.
📚References
Barth, D., & Behnke, A. (2011). The mitochondrial genome sequence of the ciliate Paramecium caudatum reveals a shift in nucleotide composition and codon usage within the genus Paramecium. BMC Genomics, 12, 272. https://doi.org/10.1186/1471-2164-12-272
Bouhouche, K., Yano, J., & Valentine, M. S. (2022). Paramecium, a model to study ciliary beating and ciliogenesis. Frontiers in Cell and Developmental Biology, 10, 847908. https://doi.org/10.3389/fcell.2022.847908
Brette, R. (2021). Integrative neuroscience of Paramecium, a "swimming neuron". eNeuro, 8(3), ENEURO.0018-21.2021. https://doi.org/10.1523/ENEURO.0018-21.2021
Catania, F., Vitali, V., Rothering, R., & Zufall, R. A. (2024). Paramecium genetics, genomics, and evolution. Annual Review of Genetics, 58, 1–25. https://doi.org/10.1146/annurev-genet-071819-104035
Fokin, S. I. (2010). Paramecium genus: biodiversity, some morphological features and the key to the main morphospecies discrimination. Protistology, 6(4), 227–235. https://www.zin.ru/journals/protistology/num6_4/fokin_227-235.pdf
Furrer, H., et al. (2024). Application of RNA interference and protein localization to study Paramecium caudatum. Communications Biology, 7, 246. https://doi.org/10.1038/s42003-024-05906-2
Gause, G. F. (1934). The Struggle for Existence. Williams and Wilkins.
Görtz, H. D., Strüder-Kypke, M., & Zeini, A. (2009). Holospora and Caedibacter: endosymbionts of Paramecium. In Endosymbionts in Paramecium (pp. 161–176). Springer.
Harumoto, T., & Miyake, A. (1991). Defensive function of trichocysts in Paramecium. Journal of Experimental Zoology, 260(1), 84–92. https://doi.org/10.1002/jez.1402600111
Heydarnejad, M. S. (2008). Survival of Paramecium caudatum at various pH values and under hypoxic condition. Pakistan Journal of Biological Sciences, 11(3), 392–397. https://doi.org/10.3923/pjbs.2008.392.397
Hori, M., & Fujishima, M. (2003). The endosymbiotic bacterium Holospora obtusa enhances heat-shock gene expression of the host Paramecium caudatum. Journal of Eukaryotic Microbiology, 50(4), 293–298. https://doi.org/10.1111/j.1550-7408.2003.tb00137.x
Komori, R., Sato, H., Harumoto, T., & Takagi, Y. (1979). Clonal death associated with the number of fissions in Paramecium caudatum. Journal of Cell Science, 41(1), 177–191. https://doi.org/10.1242/jcs.41.1.177
Krenek, S., Petzoldt, T., &325zel, T. (2012). Thermal performance curves of Paramecium caudatum: a model selection approach. European Journal of Protistology, 48(2), 124–130. https://doi.org/10.1016/j.ejop.2011.07.004
Lynn, D. H. (2010). The Ciliated Protozoa: Characterization, Classification, and Guide to the Literature (3rd ed.). Springer. https://doi.org/10.1007/978-1-4020-8239-9
McGrath, C. L., Gout, J. F., Doak, T. G., Yanagi, A., & Lynch, M. (2014). Insights into three whole-genome duplications gleaned from the Paramecium caudatum genome sequence. Genetics, 197(4), 1417–1428. https://doi.org/10.1534/genetics.114.163287
Mikami, K. (1988). Nuclear differentiation in Paramecium caudatum. Zoological Science, 5, 915–926.
Miyake, A. (1996). Defensive function of trichocysts in Paramecium against the predatory ciliate Monodinium balbiani. European Journal of Protistology, 32, 128–133. https://doi.org/10.1016/S0932-4739(96)80048-4
Patterson, D. J. (1980). Contractile vacuoles and associated structures: their organization and function. Biological Reviews, 55(1), 1–46. https://doi.org/10.1111/j.1469-185X.1980.tb00686.x
Strüder-Kypke, M. C., Wright, A.-D. G., Fokin, S. I., & Lynn, D. H. (2000). Phylogenetic relationships of the Subclass Peniculia (Oligohymenophorea, Ciliophora) inferred from small subunit rRNA gene sequences. Journal of Eukaryotic Microbiology, 47(3), 419–429. https://doi.org/10.1111/j.1550-7408.2000.tb00069.x
Uezu, T., Takei, Y., & Hiwatashi, K. (2009). Novel features of computer-simulated clonal life of Paramecium caudatum. Journal of Theoretical Biology, 259(1), 143–150. https://doi.org/10.1016/j.jtbi.2009.01.034
Valentine, M. S., & Van Houten, J. L. (2021). Using Paramecium as a model for ciliopathies. Genes, 12(10), 1493. https://doi.org/10.3390/genes12101493
Van Houten, J. L. (2023). A review for the special issue on Paramecium as a modern model organism. Microorganisms, 11(4), 890. https://doi.org/10.3390/microorganisms11040890
Wichterman, R. (2012). The Biology of Paramecium (2nd ed.). Springer. ISBN 978-1-4757-0372-6.
Yanagi, A., & Haga, N. (1998). Induction of conjugation by methyl cellulose in Paramecium caudatum. Zoological Science, 15(4), 555–559.
Gallery
2 images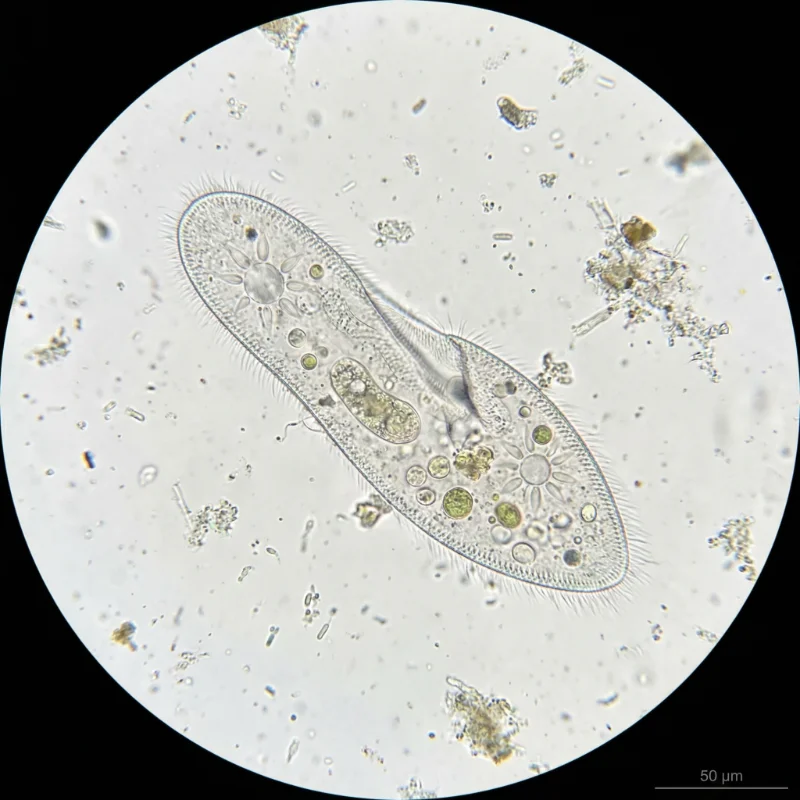 Slipper Animalcule
Slipper AnimalculeSlipper Animalcule
 Slipper Animalcule
Slipper AnimalculeSlipper Animalcule