Influenza Virus
Viruses · other
Influenzavirus A, B, C, D
Name meaning: "From Latin 'influentia' (influence), reflecting the medieval belief that the disease was caused by the astrological influence of the stars. The family name Orthomyxoviridae combines Greek 'myxa' (mucus) and 'orthos' (correct/straight), reflecting the virus's affinity for mucous membranes"
Discovery
Microorganism traits
Classification
Habitat
Classification history
1933: Smith, Andrewes, and Laidlaw first isolated human influenza virus. Subsequently, type B was identified in 1940 and type C in 1947. Type D was first identified in 2011. Classified by ICTV into four genera within the family Orthomyxoviridae. 2021-2023: ICTV mandated binomial nomenclature, resulting in species renaming (e.g., Alphainfluenzavirus influenzae)
Clinical significance
Causative agent of seasonal influenza (approximately 1 billion infections and 290,000-650,000 deaths globally per year). Capable of causing pandemics (1918 Spanish Flu, 2009 Swine Flu, etc.). Ongoing zoonotic risk from highly pathogenic avian influenza H5N1

Influenza virus (Influenzavirus) is a segmented negative-sense single-stranded RNA virus belonging to the family Orthomyxoviridae that causes acute respiratory infections in humans and a wide range of animals. Under the current taxonomy of the International Committee on Taxonomy of Viruses (ICTV), the four types of influenza — A, B, C, and D — are classified into four separate genera: Alphainfluenzavirus, Betainfluenzavirus, Gammainfluenzavirus, and Deltainfluenzavirus, respectively. Following the ICTV mandate for binomial species nomenclature beginning in 2021, the species names have been formalized as Alphainfluenzavirus influenzae, Betainfluenzavirus influenzae, Gammainfluenzavirus influenzae, and Deltainfluenzavirus influenzae (NCBI Insights, 2023).
According to the World Health Organization (WHO), approximately 1 billion seasonal influenza infections occur globally each year, of which 3 to 5 million progress to severe illness and an estimated 290,000 to 650,000 result in respiratory-related deaths (WHO, 2025). In the United States, the CDC estimated the 2024-2025 influenza season resulted in approximately 51 million influenza illnesses, 710,000 hospitalizations, and 45,000 deaths, making it a high-severity season (CDC, 2026). In developing countries, 99% of influenza-associated lower respiratory infection deaths in children under five years of age occur.
The name "influenza" derives from the Latin "influentia," reflecting the medieval belief that the disease was caused by the astrological "influence" of the stars. The family name "Orthomyxoviridae" combines the Greek words "myxa" (mucus) and "orthos" (correct, straight), reflecting the virus's tropism for mucous membranes. In 1933, Wilson Smith, Christopher Andrewes, and Patrick Laidlaw first isolated influenza virus from humans in London, England. Earlier, in 1931, Richard Shope had isolated influenza virus from swine.
1. Overview
Influenza virus is one of the most significant respiratory viral pathogens faced by humanity, posing a persistent public health threat through seasonal epidemics and irregular pandemics. In the Baltimore classification system, it belongs to Group V (negative-sense single-stranded RNA viruses), and its segmented genome provides the capacity for reassortment — the fundamental basis for pandemic potential. Seasonal influenza vaccines remain the most effective preventive measure, but the virus's continuous antigenic variation necessitates annual updates to vaccine composition.
The influenza viruses currently circulating seasonally in humans are A(H1N1)pdm09, A(H3N2), and B/Victoria lineage. The B/Yamagata lineage has not been detected globally since March 2020, likely driven to probable extinction by the non-pharmaceutical interventions implemented during the COVID-19 pandemic. Consequently, all influenza vaccines have transitioned to a trivalent formulation beginning with the 2025-2026 season (CDC, 2026; WHO, 2025). Meanwhile, highly pathogenic avian influenza A(H5N1) continues to be a cause for concern, particularly following unprecedented outbreaks in U.S. dairy cattle beginning in 2024, with sporadic human infections at the animal-human interface being reported.
2. Taxonomy and Phylogeny
The taxonomic classification of influenza viruses is as follows.
| Taxonomic Level | Name |
|---|---|
| Realm | Riboviria |
| Kingdom | Orthornavirae |
| Phylum | Negarnaviricota |
| Subphylum | Polyploviricotina |
| Class | Insthoviricetes |
| Order | Articulavirales |
| Family | Orthomyxoviridae |
| Genus | Alphainfluenzavirus (type A), Betainfluenzavirus (type B), Gammainfluenzavirus (type C), Deltainfluenzavirus (type D) |
The ICTV began mandating binomial nomenclature for all virus species in 2021, and accordingly the former "Influenza A virus" was renamed Alphainfluenzavirus influenzae, and "Influenza B virus" became Betainfluenzavirus influenzae. These changes were reflected in NCBI Taxonomy in 2023 (NCBI Insights, 2023). The family Orthomyxoviridae also includes genera beyond the four influenza virus genera, such as Thogotovirus, Quaranjavirus, and Isavirus.
Influenza A virus (Alphainfluenzavirus influenzae) is the most medically significant influenza type, capable of causing both seasonal epidemics and pandemics in humans. It is classified into subtypes based on the combination of its surface proteins hemagglutinin (HA) and neuraminidase (NA). To date, 18 HA subtypes (H1-H18) and 11 NA subtypes (N1-N11) have been identified. The major subtypes currently circulating seasonally in humans are A(H1N1) and A(H3N2). Wild waterfowl serve as the natural reservoir for influenza A viruses, and nearly all HA and NA subtype combinations have been found in avian species.
Influenza B virus (Betainfluenzavirus influenzae) primarily infects humans and is classified not by subtype but by lineage. Two lineages have been recognized — B/Victoria and B/Yamagata — although B/Yamagata has not been detected globally since March 2020 and is considered to have undergone probable extinction (Caini et al., Lancet Microbe, 2024). Non-pharmaceutical interventions during the COVID-19 pandemic were identified as the primary driver of its disappearance; while B/Victoria resurged after measures were relaxed, B/Yamagata did not recover. All influenza vaccines have consequently transitioned to trivalent formulations excluding B/Yamagata from the 2025-2026 season onward (FDA, 2025; CDC, 2026). Influenza B evolves more slowly both genetically and antigenically than influenza A and, lacking an animal reservoir, cannot undergo antigenic shift and does not cause pandemics.
Influenza C virus (Gammainfluenzavirus influenzae) infects humans and swine, causing mild upper respiratory infections or asymptomatic infections. Its genome consists of 7 segments (compared to 8 for types A and B), and it possesses a single hemagglutinin-esterase-fusion (HEF) protein that performs the functions of both HA and NA. It does not cause seasonal epidemics or pandemics and is of low public health significance.
Influenza D virus (Deltainfluenzavirus influenzae), the most recently discovered type, was first identified in 2011 and primarily infects cattle and swine. Human infection is rare, though serological evidence of exposure has been reported in livestock workers. Like influenza C, it has 7 genome segments. Its capacity to cause disease in humans has not been established.
3. Morphology and Structure
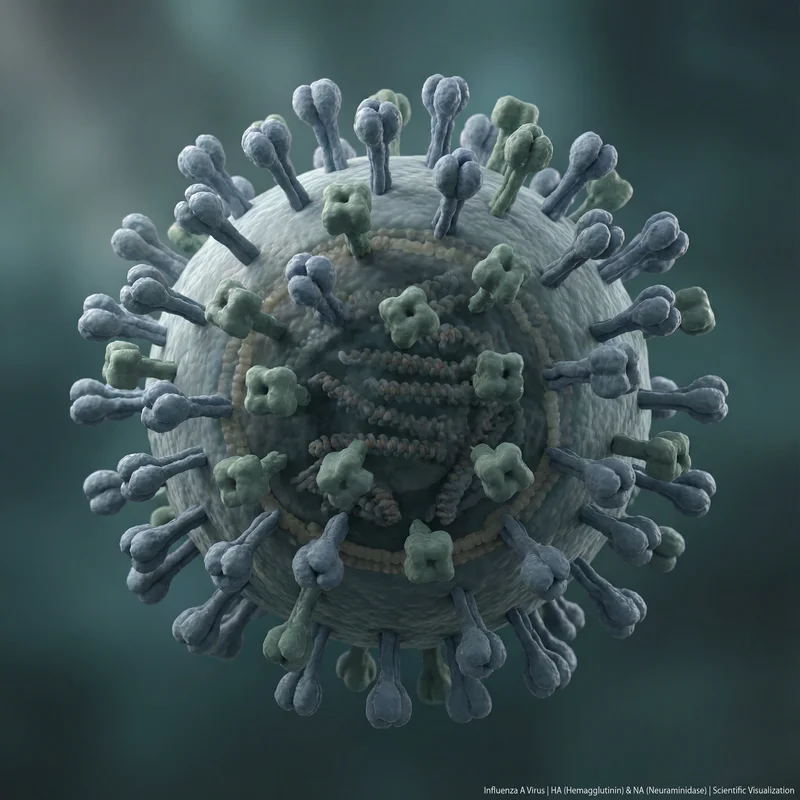
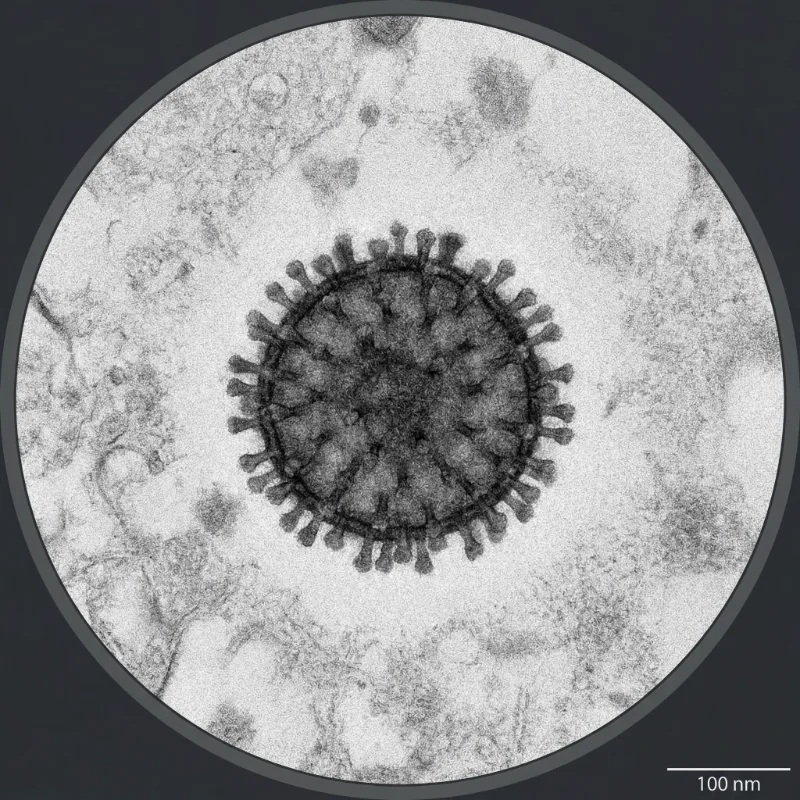
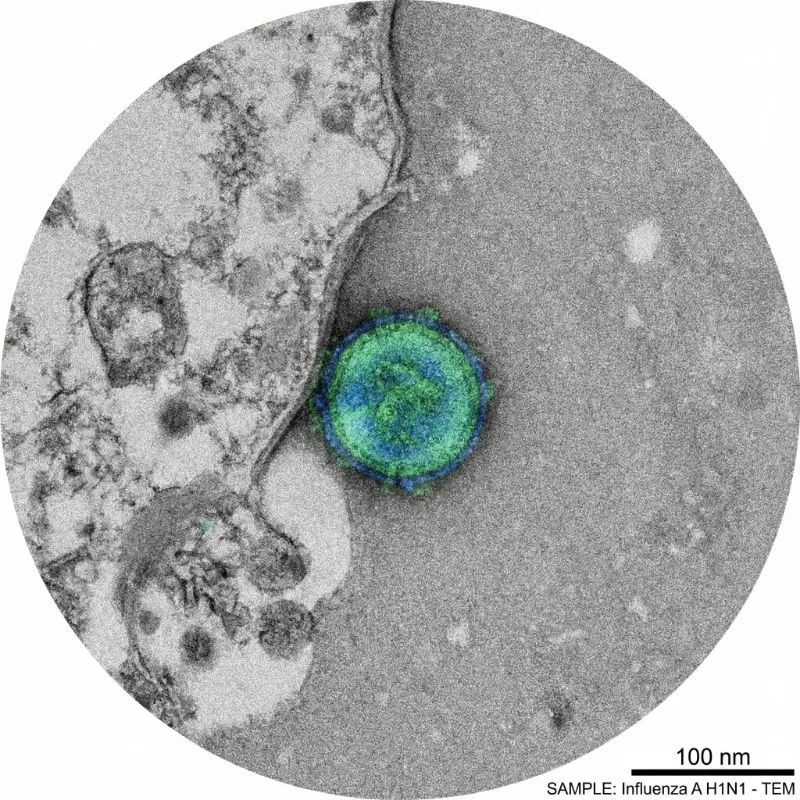
Influenza viruses are pleomorphic enveloped viruses that exist in two morphological forms: spherical and filamentous. Spherical virions have a diameter of approximately 80-120 nm, while filamentous forms have a diameter of approximately 80 nm and can extend from 300 nm to several micrometers in length. Laboratory-adapted strains are predominantly spherical, but clinical isolates more commonly exhibit filamentous morphology. The virion surface is studded with HA and NA glycoprotein spikes protruding approximately 7-14 nm, with HA outnumbering NA by a ratio of roughly 4-5:1.
Hemagglutinin (HA) is the most abundant surface glycoprotein (approximately 500 copies per virion) and exists as a homotrimer. HA initiates viral entry by binding sialic acid receptors on host cell surfaces and mediates membrane fusion upon endosomal acidification. HA consists of HA1 (receptor-binding domain) and HA2 (fusion domain), and cleavage by host proteases is essential for activation of infectivity.
Neuraminidase (NA) is a tetrameric enzymatic glycoprotein that cleaves sialic acid residues, facilitating the release of newly formed virions from host cells and promoting transit through the mucus layer. NA is the molecular target of the antiviral drugs oseltamivir (Tamiflu) and zanamivir (Relenza).
The M1 (matrix 1) protein forms a layer beneath the viral envelope, providing structural stability and playing essential roles in virion assembly and budding. The M2 protein functions as a tetrameric proton ion channel that transmits endosomal acidification into the virion interior during uncoating, facilitating the dissociation of M1 from vRNPs. The NS1 protein is a major virulence factor that antagonizes the host interferon response, while NEP (NS2) mediates the nuclear export of vRNPs.
4. Genome and Molecular Biology
Influenza A and B viruses possess a genome of 8 segments of negative-sense single-stranded RNA, while influenza C and D have 7 segments. The total genome size is approximately 13.5 kb (types A and B). Each RNA segment exists as a viral ribonucleoprotein (vRNP) complex, associated with nucleoprotein (NP) and the RNA-dependent RNA polymerase (RdRp) complex comprising PB1, PB2, and PA subunits.
The eight RNA segments of influenza A virus encode the following proteins.
| Segment | Size (nt) | Major Protein(s) | Function |
|---|---|---|---|
| 1 (PB2) | ~2,341 | PB2 | RNA polymerase subunit, cap binding |
| 2 (PB1) | ~2,341 | PB1, PB1-F2 | RNA polymerase subunit, apoptosis induction |
| 3 (PA) | ~2,233 | PA, PA-X | RNA polymerase subunit, endonuclease activity |
| 4 (HA) | ~1,778 | HA (hemagglutinin) | Receptor binding, membrane fusion |
| 5 (NP) | ~1,565 | NP (nucleoprotein) | vRNA binding, vRNP structural component |
| 6 (NA) | ~1,413 | NA (neuraminidase) | Sialic acid cleavage, virion release |
| 7 (M) | ~1,027 | M1, M2 | Matrix protein, ion channel |
| 8 (NS) | ~890 | NS1, NEP/NS2 | Interferon antagonist, vRNP nuclear export |
The influenza virus replication cycle consists of the following stages. First, during the attachment phase, HA binds sialic acid receptors on the host cell surface. Human-adapted viruses preferentially bind alpha-2,6-linked sialic acid, while avian-adapted viruses prefer alpha-2,3-linked sialic acid. Second, the virus is internalized via receptor-mediated endocytosis. Third, endosomal acidification (pH ~5) triggers a conformational change in HA that initiates fusion between the viral envelope and endosomal membrane, while simultaneously the M2 ion channel allows proton influx, releasing vRNPs. Fourth, vRNPs are transported to the nucleus, where viral RNA transcription and replication occur — a distinctive feature of influenza among RNA viruses, most of which replicate in the cytoplasm. Fifth, viral mRNAs are translated in the cytoplasm, and new vRNPs are assembled and exported from the nucleus. Finally, viral proteins and vRNPs are assembled at the plasma membrane, and NA cleaves sialic acid to allow new virions to bud from the cell.
The mutation rate of influenza virus is approximately 2.3 x 10-5 substitutions per nucleotide per replication cycle, which is high among RNA viruses. This is because the RNA-dependent RNA polymerase lacks proofreading capability. The combination of a high mutation rate and immune selection pressure enables rapid antigenic variation. Additionally, the segmented genome allows reassortment when different viruses co-infect the same cell, potentially generating entirely novel viral combinations.
5. Antigenic Variation
Antigenic drift is the gradual accumulation of point mutations in the HA and NA genes, resulting in progressive antigenic change. Under immune selection pressure, mutations accumulate at antibody-binding sites (antigenic epitopes), and variants capable of evading pre-existing immunity are selected. This continuous drift necessitates annual review and updating of seasonal influenza vaccine composition. The H3N2 subtype undergoes antigenic drift more rapidly than H1N1, and influenza B evolves more slowly than influenza A (Petrova & Russell, Nature Reviews Microbiology, 2018).
Antigenic shift is the abrupt antigenic change that occurs when different influenza A viruses co-infect the same host cell and undergo reassortment of genome segments. When a virus with a novel HA and/or NA subtype emerges, the majority of the population is immunologically naive, creating conditions for a pandemic. All influenza pandemics of the 20th and 21st centuries resulted from antigenic shift. Influenza B, lacking an animal reservoir, cannot undergo antigenic shift and does not cause pandemics.
6. Pathogenesis and Clinical Features
Typical influenza symptoms include sudden onset of fever (38-40°C), chills, headache, myalgia, arthralgia, and fatigue, accompanied by respiratory symptoms such as dry cough, sore throat, and nasal congestion. However, only approximately 50% of infected individuals exhibit these classic symptoms. In children, gastrointestinal symptoms including nausea, vomiting, and diarrhea are more common. In most healthy adults, symptoms resolve spontaneously within 7-10 days, though cough and fatigue may persist for several weeks.
The incubation period averages approximately 2 days (range: 1-4 days). Infected individuals begin shedding virus approximately 1 day before symptom onset and remain infectious for 5-7 days after symptom onset. Children and immunocompromised individuals may shed virus for longer periods. Infectiousness is highest during the first 2-3 days following symptom onset.
Severe complications include viral pneumonia, secondary bacterial pneumonia (particularly Streptococcus pneumoniae and Staphylococcus aureus), acute respiratory distress syndrome (ARDS), myocarditis, encephalitis, rhabdomyolysis, and multi-organ failure. High-risk groups include adults aged 65 and older, children under 5 (especially under 2), pregnant women, individuals with chronic pulmonary or cardiovascular disease, diabetes, immunocompromised patients, and obese individuals (BMI >= 40).
Reverse transcription polymerase chain reaction (RT-PCR) is the gold standard for influenza diagnosis, offering high sensitivity and specificity with the ability to distinguish virus type (A, B) and subtype (H1, H3, N1, N2, etc.). Multiplex RT-PCR can simultaneously detect influenza and other respiratory viruses (RSV, SARS-CoV-2, etc.). Rapid influenza diagnostic tests (RIDTs) provide results within 15-30 minutes with sensitivity of 50-70% and specificity exceeding 90%. Virus culture is used for detailed antigenic characterization and antiviral susceptibility testing, and is essential for surveillance and vaccine strain selection.
7. Epidemiology and Transmission
Seasonal influenza epidemics occur primarily during winter months in temperate regions (November-March in the Northern Hemisphere, May-September in the Southern Hemisphere), while year-round transmission occurs in tropical regions. WHO estimates approximately 1 billion annual infections, 3-5 million severe cases, and 290,000-650,000 respiratory-associated deaths globally (WHO, 2025). In the United States, the 2024-2025 season was a high-severity season with an estimated 51 million influenza illnesses, 23 million medical visits, 710,000 hospitalizations, and 45,000 deaths (CDC, 2026).
The major influenza pandemics since the 20th century are summarized below.
| Pandemic | Year | Causative Virus | Origin | Estimated Deaths |
|---|---|---|---|---|
| Spanish Flu | 1918-1919 | A(H1N1) | Uncertain (probable avian origin) | 50-100 million |
| Asian Flu | 1957-1958 | A(H2N2) | Avian-human reassortment | 1-2 million |
| Hong Kong Flu | 1968-1969 | A(H3N2) | Avian-human reassortment | 1 million |
| Russian Flu | 1977-1978 | A(H1N1) | Probable laboratory release | Unknown |
| Swine Flu | 2009-2010 | A(H1N1)pdm09 | Swine-avian-human reassortment | 150,000-575,000 |
The 1918 Spanish Flu remains one of the deadliest pandemics in human history, infecting approximately one-third of the world's population and killing an estimated 50 million or more people. Unusually, it caused high mortality among healthy adults aged 20-40, which is believed to be related to an exaggerated immune response ("cytokine storm") (Taubenberger & Morens, Emerging Infectious Diseases, 2006).
Highly pathogenic avian influenza A(H5N1) was first identified in humans in Hong Kong in 1997. According to WHO cumulative data as of December 19, 2025, a total of 993 confirmed human cases have been reported worldwide since 2003, with 477 deaths, yielding a case fatality rate of approximately 48% (WHO, December 2025). Sustained human-to-human transmission has not been documented. However, beginning in March 2024, unprecedented outbreaks of H5N1 were identified in U.S. dairy cattle herds, with over 1,070 herd infections confirmed across more than 16 states by May 2025, along with 71 confirmed human infections in the United States (CDC, 2025). Active surveillance and control measures subsequently reduced these outbreaks substantially, but widespread circulation in wild bird populations persists worldwide.
8. Immunity, Treatment, and Prevention
FDA-approved influenza antiviral drugs include neuraminidase inhibitors (oseltamivir, zanamivir, peramivir) and the cap-dependent endonuclease inhibitor baloxavir marboxil. Oseltamivir (Tamiflu), the most widely used oral agent, reduces symptom duration by approximately 1 day and lowers the risk of complications when administered within 48 hours of symptom onset. Zanamivir (Relenza) is an inhaled formulation, peramivir (Rapivab) is given intravenously, and baloxavir (Xofluza) offers the advantage of a single oral dose treatment.
| Drug | Brand Name | Mechanism | Route | Regimen |
|---|---|---|---|---|
| Oseltamivir | Tamiflu | NA inhibitor | Oral | 75 mg twice daily for 5 days |
| Zanamivir | Relenza | NA inhibitor | Inhaled | 10 mg twice daily for 5 days |
| Peramivir | Rapivab | NA inhibitor | Intravenous | 600 mg single dose |
| Baloxavir marboxil | Xofluza | Endonuclease inhibitor | Oral | Weight-based single dose |
The M2 ion channel inhibitors amantadine and rimantadine, once used against influenza A, are no longer recommended because virtually all currently circulating influenza A viruses carry the M2 resistance mutation S31N.
Seasonal influenza vaccination is the most effective preventive measure. Vaccine types include inactivated influenza vaccine (IIV), recombinant influenza vaccine (RIV), and live attenuated influenza vaccine (LAIV). Beginning with the 2025-2026 season, all influenza vaccines contain three antigens — A(H1N1), A(H3N2), and B/Victoria — in a trivalent formulation (FDA, 2025; CDC, 2026). Vaccine effectiveness varies depending on the antigenic match between vaccine strains and circulating viruses, generally providing 40-60% protection against symptomatic infection. Effectiveness against severe disease, hospitalization, and death is higher.
| Vaccine Type | Abbreviation | Characteristics | Eligible Ages |
|---|---|---|---|
| Inactivated (standard dose) | IIV-SD | Injection, most widely used | 6 months and older |
| Inactivated (high dose) | IIV-HD | For ages 65+, 4x antigen | 65 and older |
| Inactivated (adjuvanted) | aIIV | MF59 adjuvant | 65 and older |
| Recombinant | RIV | Egg-free, 3x antigen | 18 and older |
| Live attenuated | LAIV | Nasal spray | 2-49 years |
WHO and national health authorities recommend annual influenza vaccination for high-risk groups including adults 65 and older, children 6 months to 5 years, pregnant women, individuals with chronic medical conditions, healthcare workers, and residents and staff of long-term care facilities.
Development of a universal influenza vaccine is actively underway. Current vaccines target the variable head region of HA and require annual updates, whereas universal vaccines targeting conserved antigens such as the HA stalk, M2e, or neuraminidase could provide broad, durable protection. As of February 2026, CIDRAP's Universal Influenza Vaccine Technology Landscape documents numerous candidate vaccines in clinical trials, and mRNA-based seasonal influenza vaccines (such as Moderna's mRNA-1010) have reported Phase 3 clinical trial results (CIDRAP, 2026). In December 2025, WHO released updated Preferred Product Characteristics to guide the development of next-generation influenza vaccines (WHO, 2026).
Hand hygiene, respiratory etiquette, mask wearing, avoiding contact with symptomatic individuals, and ventilation are effective in reducing transmission. Symptomatic individuals are recommended to self-isolate until at least 24 hours after fever resolves.
9. Natural Hosts and Animal Infections
Wild waterfowl (ducks, geese, swans) are the principal natural reservoir hosts of influenza A viruses, and nearly all HA (18) and NA (11) subtype combinations have been identified in these species. Most avian influenza viruses cause asymptomatic intestinal infections in wild birds and are transmitted via the fecal-oral route. Migratory bird flyways play a critical role in the long-range geographic dissemination of the virus.
Domestic poultry (chickens, turkeys) are highly susceptible to avian influenza. Low pathogenicity avian influenza (LPAI) causes mild symptoms, but highly pathogenic avian influenza (HPAI, particularly certain H5 and H7 subtypes) causes acute systemic infection with mortality approaching 100% in poultry, inflicting massive economic losses.
Swine are an important host for influenza A viruses, with H1N1, H1N2, and H3N2 as the major circulating subtypes. Pigs express both alpha-2,3- and alpha-2,6-linked sialic acid receptors, enabling simultaneous infection by both avian and human viruses. This makes swine a "mixing vessel" for reassortment, a critical pathway for the emergence of novel pandemic viruses. The 2009 H1N1 pandemic virus was the product of complex reassortment involving gene segments from swine, avian, and human influenza viruses (Neumann et al., Nature, 2009).
Influenza A virus infections have also been reported in horses (H3N8, H7N7), dogs (H3N8, H3N2), cats, and marine mammals (seals, whales). Beginning in March 2024, unprecedented outbreaks of H5N1 were confirmed in U.S. dairy cattle herds, representing the first known involvement of cattle as significant hosts. The initial outbreaks were caused by the B3.13 genotype, and a novel D1.1 genotype was subsequently identified in dairy cattle in early 2025 (USDA, 2025). Although active control measures substantially reduced these outbreaks, the event underscored the unpredictable host-range expansion capacity of influenza viruses.
10. Research History and Scientific Significance
Influenza virus research has been central to the advancement of modern virology. In 1931, Richard Shope isolated influenza virus from swine, marking the beginning of animal influenza research. In 1933, Wilson Smith, Christopher Andrewes, and Patrick Laidlaw achieved the first isolation of influenza virus from humans (Smith et al., The Lancet, 1933). In the 1940s, Thomas Francis Jr. and Jonas Salk developed the first inactivated influenza vaccines, which have been continuously improved since.
In 2005, the research team led by Jeffery Taubenberger fully reconstructed the genome of the 1918 Spanish Flu virus from tissue preserved in permafrost and formalin-fixed specimens, elucidating the genetic features responsible for its extreme virulence. The development of reverse genetics technology has revolutionized influenza virus research and vaccine development.
Current major research directions include development of universal influenza vaccines, mRNA platform-based influenza vaccines, elucidation of cross-species transmission mechanisms of avian influenza, investigation of post-influenza long-term sequelae ("long flu"), surveillance of antiviral drug resistance, and understanding the mechanisms underlying seasonal influenza epidemics.
11. Comparison of Influenza Types
| Feature | Influenza A | Influenza B | Influenza C | Influenza D |
|---|---|---|---|---|
| Genome segments | 8 | 8 | 7 | 7 |
| Host range | Humans, birds, swine, horses, etc. | Humans (marine mammals rarely) | Humans, swine | Cattle, swine |
| Subtype/lineage | HA, NA subtypes | Lineages (Victoria, Yamagata) | N/A | N/A |
| Antigenic shift | Occurs | Does not occur | Does not occur | Does not occur |
| Pandemic potential | High | None | None | None |
| Disease severity | Moderate to severe | Mild to moderate | Mild | Not established in humans |
| Seasonal epidemics | Major cause | Major cause | Rare | N/A |
| Included in vaccine | Yes | Yes | No | No |
Influenza A virus, with its broad host range and capacity for antigenic shift, poses the highest pandemic potential and remains the primary focus of global surveillance. Influenza B, although confined to predominantly human circulation, still causes substantial disease burden particularly in children and the elderly.
12. Unresolved Questions
First, the timeline and achievable efficacy of universal influenza vaccines remain uncertain. Various approaches targeting conserved antigens including the HA stalk, M2e, and neuraminidase are in clinical trials, but further research is needed to demonstrate broad and durable protective efficacy.
Second, the precise genetic changes required for H5N1 and other avian influenza viruses to acquire efficient human-to-human transmission capability have not been fully elucidated. The 2024-2025 U.S. dairy cattle outbreaks provided new data on mammalian adaptation, but significant uncertainty persists in pandemic risk assessment.
Third, the exact mechanisms underlying the seasonal epidemic patterns of influenza remain debated. Temperature, humidity, ultraviolet radiation, human behavioral patterns, and population immunity status are all believed to contribute.
Fourth, monitoring and responding to the emergence and spread of antiviral resistance remains an ongoing challenge. Treatment-emergent resistance has been reported with baloxavir, requiring continued vigilance.
Fifth, research on post-influenza long-term sequelae ("long flu") is in its early stages. Similar to long COVID following SARS-CoV-2 infection, persistent fatigue, cognitive impairment, and other symptoms have been reported following influenza infection, but their frequency and mechanisms remain poorly characterized.
13. References
Smith, W., Andrewes, C.H., & Laidlaw, P.P. (1933). A virus obtained from influenza patients. The Lancet, 222(5732), 66-68. https://doi.org/10.1016/S0140-6736(00)78541-2
Webster, R.G., Bean, W.J., Gorman, O.T., et al. (1992). Evolution and ecology of influenza A viruses. Microbiological Reviews, 56(1), 152-179. https://doi.org/10.1128/mr.56.1.152-179.1992
Taubenberger, J.K., & Morens, D.M. (2006). 1918 Influenza: the mother of all pandemics. Emerging Infectious Diseases, 12(1), 15-22. https://doi.org/10.3201/eid1201.050979
Neumann, G., Noda, T., & Kawaoka, Y. (2009). Emergence and pandemic potential of swine-origin H1N1 influenza virus. Nature, 459(7249), 931-939. https://doi.org/10.1038/nature08157
Gamblin, S.J., & Skehel, J.J. (2010). Influenza hemagglutinin and neuraminidase membrane glycoproteins. Journal of Biological Chemistry, 285(37), 28403-28409. https://doi.org/10.1074/jbc.R110.129809
Krammer, F., Smith, G.J.D., Fouchier, R.A.M., et al. (2018). Influenza. Nature Reviews Disease Primers, 4, 3. https://doi.org/10.1038/s41572-018-0002-y
Petrova, V.N., & Russell, C.A. (2018). The evolution of seasonal influenza viruses. Nature Reviews Microbiology, 16(1), 47-60. https://doi.org/10.1038/nrmicro.2017.118
Long, J.S., Mistry, B., Haslam, S.M., & Barclay, W.S. (2019). Host and viral determinants of influenza A virus species specificity. Nature Reviews Microbiology, 17(2), 67-81. https://doi.org/10.1038/s41579-018-0115-z
Morens, D.M., Taubenberger, J.K., & Fauci, A.S. (2009). The persistent legacy of the 1918 influenza virus. New England Journal of Medicine, 361(3), 225-229. https://doi.org/10.1056/NEJMp0904819
Caini, S., Meijer, A., Nunes, M.C., et al. (2024). Probable extinction of influenza B/Yamagata and its public health implications. The Lancet Microbe, 5(7), e614-e622. https://doi.org/10.1016/S2666-5247(24)00066-1
World Health Organization (WHO). (2025). Influenza (Seasonal). https://www.who.int/news-room/fact-sheets/detail/influenza-(seasonal)
World Health Organization (WHO). (2025). Cumulative number of confirmed human cases for avian influenza A(H5N1) reported to WHO, 2003-2025, 19 December 2025. https://www.who.int/publications/m/item/cumulative-number-of-confirmed-human-cases-for-avian-influenza-a(h5n1)-reported-to-who--2003-2025--19-december-2025
Centers for Disease Control and Prevention (CDC). (2026). 2024-2025 Influenza Season Summary. https://www.cdc.gov/flu-burden/php/data-vis-vac/2024-2025-prevented.html
Centers for Disease Control and Prevention (CDC). (2026). 2025-2026 Flu Season. https://www.cdc.gov/flu/season/2025-2026.html
U.S. Food and Drug Administration (FDA). (2025). Influenza Vaccine Composition for the 2025-2026 U.S. Influenza Season. https://www.fda.gov/vaccines-blood-biologics/influenza-vaccine-composition-2025-2026-us-influenza-season
NCBI Insights. (2023). Upcoming changes to influenza virus names in NCBI Taxonomy. https://ncbiinsights.ncbi.nlm.nih.gov/2023/02/21/influenza-virus-ncbi-taxonomy/
Bouvier, N.M., & Palese, P. (2008). The biology of influenza viruses. Vaccine, 26(Suppl 4), D49-D53. https://doi.org/10.1016/j.vaccine.2008.07.039
Plotkin, S.A., Orenstein, W.A., Offit, P.A., & Edwards, K.M. (2018). Plotkin's Vaccines (7th ed.). Elsevier.
Yoon, S.W., Webby, R.J., & Webster, R.G. (2014). Evolution and ecology of influenza A viruses. Current Topics in Microbiology and Immunology, 385, 359-375. https://doi.org/10.1007/82_2014_396
CIDRAP. (2026). Universal Influenza Vaccine Technology Landscape. https://ivr.cidrap.umn.edu/
Fun Facts
Influenza virus is one of the rare RNA viruses that replicates inside the host cell nucleus. While most RNA viruses replicate in the cytoplasm, influenza conducts its transcription and replication within the nucleus — a strategy that allows it to exploit the host cell's splicing machinery to produce two different proteins from the M and NS gene segments.
Wild ducks and geese serve as the natural reservoir for influenza A viruses, harboring nearly all known combinations of the 18 HA and 11 NA subtypes. Remarkably, in wild waterfowl, influenza viruses replicate primarily in the intestine, causing asymptomatic infections and being shed through feces into water, which serves as the transmission route to other birds.
The genome of the 1918 Spanish Flu virus was fully reconstructed in 2005 from tissue preserved in permafrost and formalin-fixed specimens of victims. This research revealed the genetic features contributing to the virus's extreme virulence and suggested that the avian-origin virus may have adapted directly to humans without an intermediate host.
Pigs express both alpha-2,3- and alpha-2,6-linked sialic acid receptors, enabling them to be simultaneously infected by both avian and human influenza viruses. This makes swine a "mixing vessel" for genetic reassortment that can generate novel pandemic viruses — the 2009 H1N1 pandemic virus being a prime example.
The influenza virus neuraminidase (NA) enzyme cleaves sialic acid on host cell surfaces and in mucus to facilitate the release of newly formed virions. Antiviral drugs such as oseltamivir (Tamiflu) and zanamivir (Relenza) work by inhibiting NA, thereby blocking viral release and limiting the spread of infection.
The M2 protein of influenza A virus functions as a proton ion channel and was formerly the target of the antiviral drugs amantadine and rimantadine. However, virtually all currently circulating influenza A viruses carry the S31N resistance mutation in M2, rendering these drugs ineffective and no longer recommended for use.
The composition of seasonal influenza vaccines is determined through WHO's Global Influenza Surveillance and Response System (GISRS). Virus specimens collected by National Influenza Centres in over 143 countries are analyzed at 6 WHO Collaborating Centres, which inform the vaccine strain recommendations issued every February (Northern Hemisphere) and September (Southern Hemisphere).
Influenza viruses are pleomorphic, with laboratory-adapted strains being mostly spherical (about 100 nm diameter), but clinical isolates more often displaying a filamentous morphology that can reach several micrometers in length. Interestingly, the virion shape is determined by the host cell type and specific amino acid residues in the M1 matrix protein.
The 2009 pandemic virus A(H1N1)pdm09 was the product of extraordinarily complex reassortment, combining gene segments derived from North American swine influenza (a triple reassortant virus itself), Eurasian avian-lineage swine influenza, and human H3N2 influenza viruses.
The B/Yamagata lineage of influenza B virus has not been detected worldwide since March 2020, likely driven to extinction by non-pharmaceutical interventions during the COVID-19 pandemic. As a result, all influenza vaccines transitioned to a trivalent formulation containing only B/Victoria starting with the 2025-2026 season.
In March 2024, highly pathogenic avian influenza H5N1 was confirmed for the first time ever in U.S. dairy cattle — a species previously not considered a significant influenza host. This unprecedented event demonstrated the unpredictable host-range expansion capability of influenza viruses and raised new concerns about mammalian adaptation pathways.
Influenza virus employs a unique strategy called 'cap-snatching' to synthesize its mRNA. The viral polymerase steals a short capped RNA fragment (10-13 nucleotides) from host cell pre-mRNAs and uses it as a primer to initiate transcription of its own messenger RNA — a molecular theft that also disrupts host gene expression.
FAQ
Although both are respiratory viral infections, influenza and the common cold differ in causative agent and symptom severity. Influenza is caused by influenza viruses and is characterized by sudden onset of high fever (38-40°C), severe myalgia, headache, and extreme fatigue. The common cold is typically caused by rhinoviruses or seasonal coronaviruses, with predominant symptoms of runny nose, nasal congestion, sneezing, and mild sore throat; fever is rare or mild. Influenza can lead to serious complications such as pneumonia, and both vaccines and antiviral drugs are available, whereas there is no specific treatment for the common cold.
The influenza vaccine does not completely prevent symptomatic infection. Vaccine effectiveness varies depending on how well the vaccine strains match circulating viruses and generally provides 40-60% protection against symptomatic infection. However, vaccination significantly reduces the risk of severe complications, hospitalization, and death even if infection does occur. The benefits are especially pronounced in high-risk groups such as the elderly, those with chronic medical conditions, and young children. Additionally, inactivated vaccines use killed virus and cannot cause the flu.
Influenza viruses continuously change through antigenic drift — the gradual accumulation of point mutations in the HA and NA surface proteins that allow new variants to evade existing immunity. Accordingly, WHO reviews and recommends vaccine composition every February (for the Northern Hemisphere) and September (for the Southern Hemisphere), selecting the virus strains most likely to circulate in the upcoming season. Additionally, vaccine-induced immunity wanes over time, making annual vaccination recommended for optimal protection.
Avian influenza viruses, including H5N1, can infect humans, but sustained human-to-human transmission has not been documented to date. Human infections primarily result from direct exposure to infected poultry, contaminated environments, or infected dairy cattle. According to WHO cumulative data through December 2025, 993 confirmed human H5N1 cases have been reported globally since 2003, with 477 deaths, yielding a case fatality rate of approximately 48%. If the virus were to acquire the ability to transmit efficiently between humans, a severe pandemic could result, which is why continuous global surveillance and pandemic preparedness efforts are ongoing.
Oseltamivir (Tamiflu) and other influenza antivirals are most effective when administered within 48 hours of symptom onset. Early treatment can shorten symptom duration by approximately one day and reduce the risk of complications. However, even beyond 48 hours, treatment is still recommended for hospitalized patients, those with severe illness, and high-risk individuals. Not all influenza patients require antiviral treatment — mild infections in otherwise healthy adults typically resolve with supportive care. Antiviral treatment is particularly recommended for high-risk groups, severe cases, and hospitalized patients.
While symptoms of influenza A and B are similar, there are several important differences. Type A infects a wide range of hosts (humans, birds, swine, horses, etc.), is classified by HA/NA subtypes, and can cause pandemics through antigenic shift. Type B infects primarily humans, is classified by lineage (Victoria/Yamagata), lacks antigenic shift capability, and does not cause pandemics. Type A generally tends to cause more severe disease, but type B can also cause serious illness, particularly in children and the elderly. Both types are included in seasonal influenza vaccines.
Influenza-infected individuals begin shedding virus approximately 1 day before symptom onset and remain contagious for 5-7 days after symptoms appear. Infectiousness is highest during the first 2-3 days after symptom onset. Children and immunocompromised individuals may shed virus for 10 or more days. Symptomatic patients are therefore advised to self-isolate until at least 24 hours after fever resolves and to avoid close contact with others.
The 1918 Spanish Flu was deadly due to a combination of factors. First, the H1N1 virus carried entirely novel antigens to which most of the population had no pre-existing immunity. Second, specific genetic features of the virus triggered an exaggerated immune response ("cytokine storm") that caused high mortality among healthy young adults aged 20-40. Third, antibiotics were not yet available, making secondary bacterial pneumonia untreatable. Fourth, troop movements and overcrowded conditions during World War I accelerated transmission. An estimated one-third of the world's population was infected, and 50-100 million people died, making it one of the deadliest pandemics in human history.
The B/Yamagata lineage of influenza B virus has not been detected in global surveillance systems since March 2020 and is considered to have undergone probable extinction. Non-pharmaceutical interventions during the COVID-19 pandemic (masking, social distancing, travel restrictions) are identified as the primary cause. While the B/Victoria lineage resurged once these measures were relaxed, B/Yamagata did not recover. A 2024 systematic review published in The Lancet Microbe confirmed these findings, and consequently, all influenza vaccines have excluded the B/Yamagata antigen beginning with the 2025-2026 season.
According to CDC estimates, the 2024-2025 U.S. influenza season was a high-severity season. An estimated 51 million influenza-related illnesses, 23 million medical visits, 710,000 hospitalizations, and 45,000 deaths occurred. This represented one of the highest hospitalization rates recorded since the 2010-2011 season. Adults aged 75 and older bore the highest hospitalization burden at approximately 599 per 100,000 population.
Gallery
4 images Influenza Virus
Influenza VirusInfluenza Virus
 Influenza Virus
Influenza VirusInfluenza Virus
 Influenza Virus
Influenza VirusInfluenza Virus
 Influenza Virus
Influenza VirusInfluenza Virus